Introduction
In this page, we explore the relationship between promoter methylation and gene expression by directly visualizing real data from TCGA.
It is generally believed that promoter methylation suppresses gene expression.
But to what extent does this relationship actually hold?
When we examine real data, several aspects emerge that cannot be fully captured by such a simple explanation.
Let’s take a look at the data.
Using Subio Platform, TCGA RNA-Seq data and DNA methylation array data can be easily imported and compared side by side (see tutorials for importing TCGA RNA-Seq and DNA methylation data).
Here, using TCGA-LIHC (Liver Hepatocellular Carcinoma) as an example, we examine the relationship between methylation and gene expression by directly observing the data. Specifically, for each gene measured by RNA-Seq, methylation sites located within 4 kbp of the transcription start site (TSS) are mapped.
This DNA methylation data was measured using the Illumina Infinium HumanMethylation450 BeadChip. This array is designed to densely cover methylation sites near the TSS (within 200 bp and 1500 bp upstream) and CpG islands. Therefore, methylation sites are not uniformly distributed across the genome.
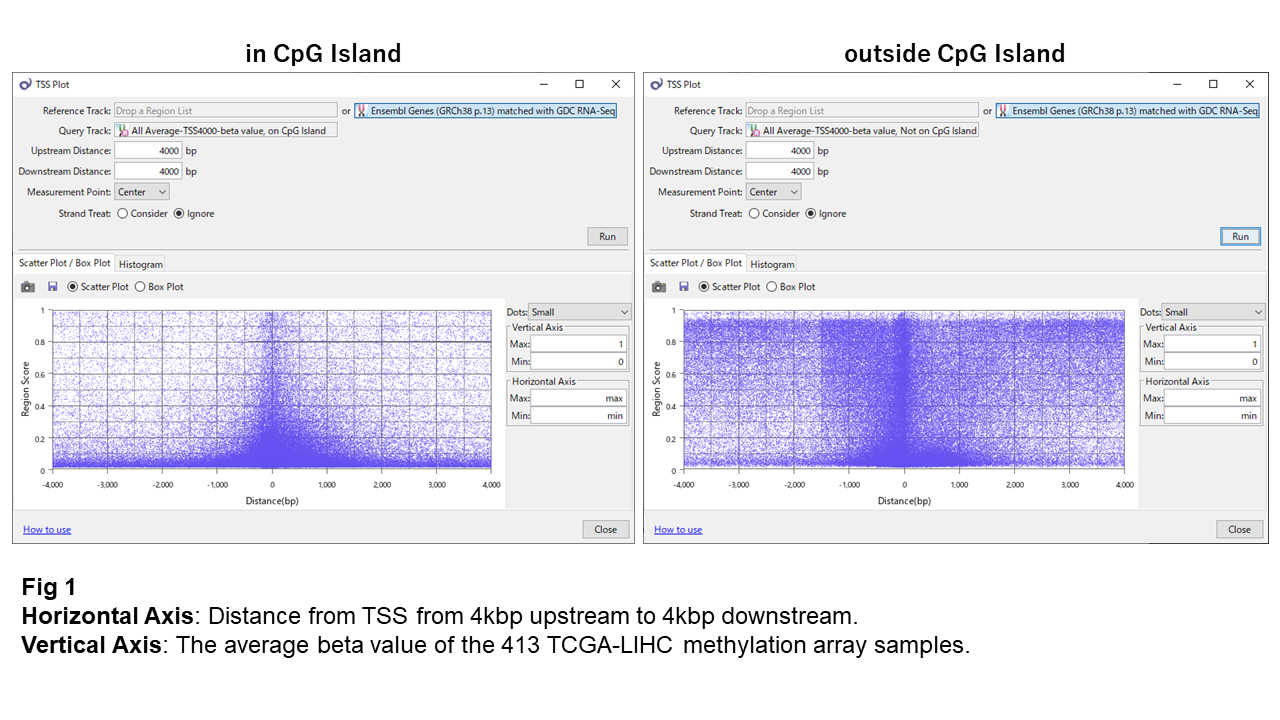
When plotting the distribution of beta values (a measure of methylation ranging from 0 to 1) around the TSS, points appear densely clustered near the TSS (Fig. 1). At first glance, this may suggest that methylation sites are concentrated near the TSS, but this pattern largely reflects the array design rather than biological reality.
Overview of TCGA-LIHC Methylation Data
Fig. 1 shows a TSS plot of beta values. Many CpG island sites are in a hypomethylated state, while some sites near the TSS show high methylation.
Outside CpG islands, most sites appear highly methylated overall. However, within ~1–1.5 kbp of the TSS, many sites are also hypomethylated.
In Fig. 2, changes in beta values between normal and tumor samples are shown. Positive values indicate increased methylation in tumors, while negative values indicate demethylation.
Within CpG islands, beta values remain relatively stable, suggesting tightly maintained methylation levels. Near the TSS, however, some sites show variation, mostly toward increased methylation.
Outside CpG islands, variation is clearly larger, and changes are strongly biased toward demethylation. Only a small fraction of sites near the TSS show increased methylation.
These observations suggest that CpG islands and non-CpG island regions may be governed by different regulatory mechanisms.
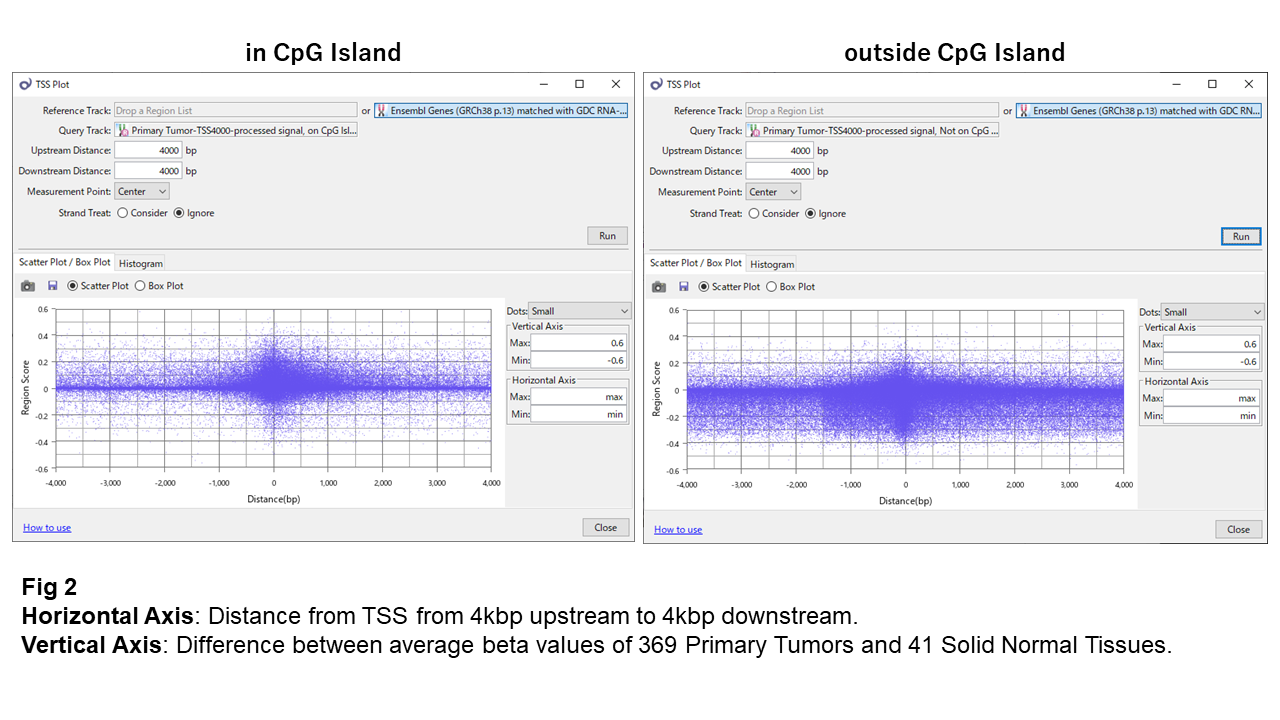
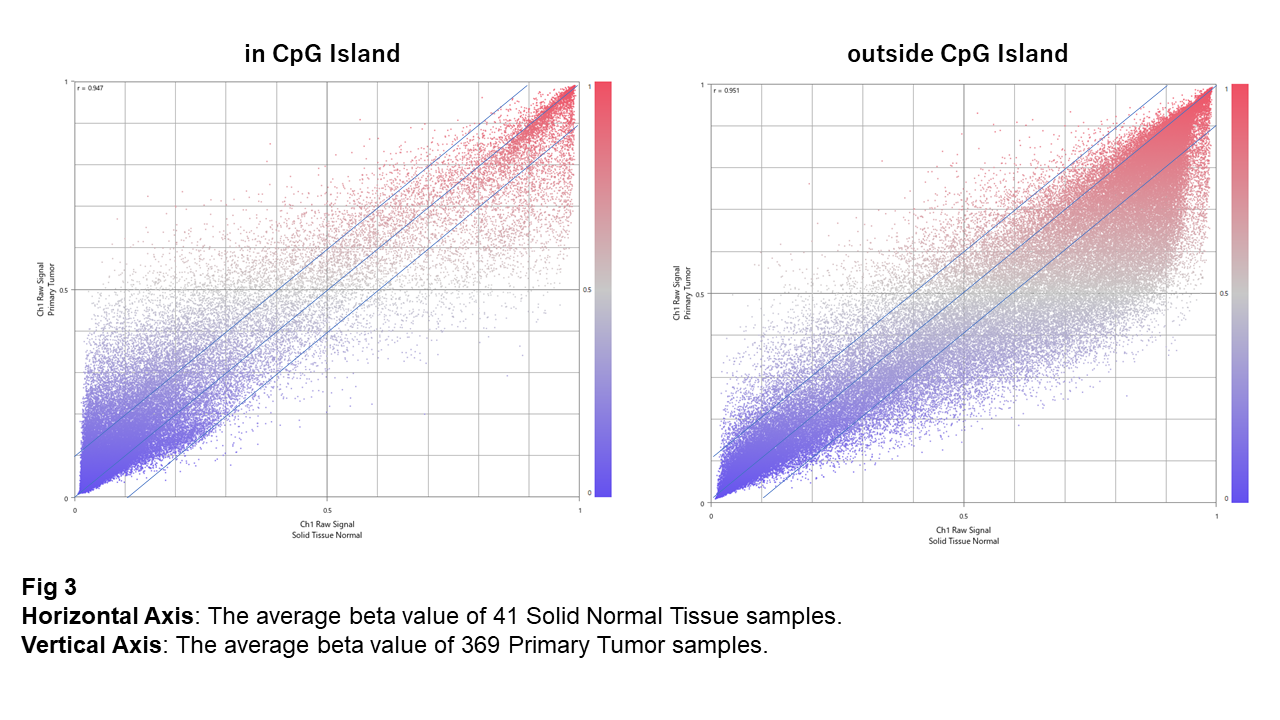
Fig. 3 is a scatter plot comparing the average beta values between Normal (x-axis) and Tumor (y-axis). Looking at the plot, within CpG islands, some sites that are originally in a hypomethylated state appear to become more methylated in tumors.
In contrast, outside CpG islands, a substantial number of sites that are originally highly methylated show changes toward demethylation in tumors.
Thus, it can be seen that the direction of change in tumors is roughly opposite between CpG islands and non-CpG island regions.
Correlation Between Methylation and Expression Patterns
Now we move to the main question: how do these methylation changes relate to gene expression?
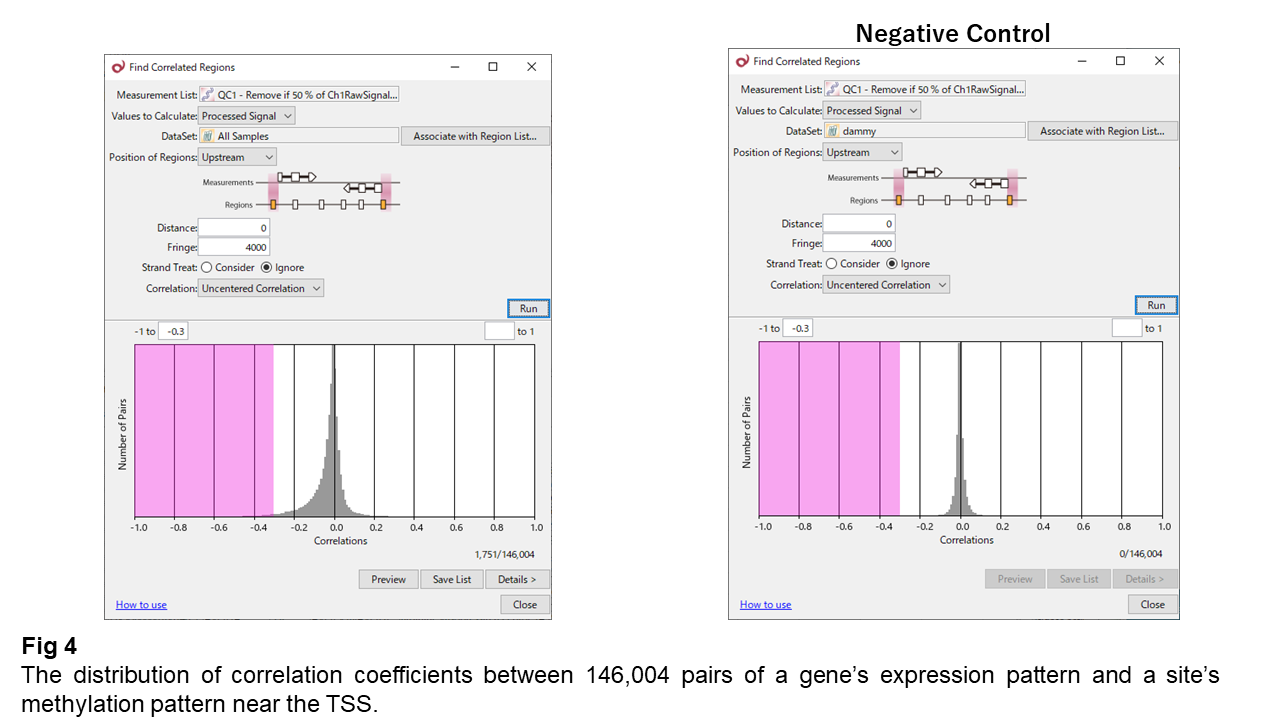
Fig. 4 shows the distribution of correlation coefficients between methylation changes and expression changes.
Most correlations cluster around zero, indicating that strong inverse correlation is generally absent. While one might expect a clear negative correlation based on textbook knowledge, the data show that such a simple relationship does not hold.
However, the distribution is slightly skewed toward negative values, indicating the presence of weak inverse correlations.
When sample labels are randomly shuffled (negative control), this bias disappears, suggesting that the observed skew is not due to chance but reflects a statistically meaningful pattern.
Spatial Distribution of Correlations
Fig. 5 shows that sites with inverse correlation are concentrated within ~300 bp of the TSS and become rare further away.
Although more such sites appear within CpG islands than outside, this is due to probe density. When comparing proportions, both are nearly identical (~1.4% vs. 1.3%).
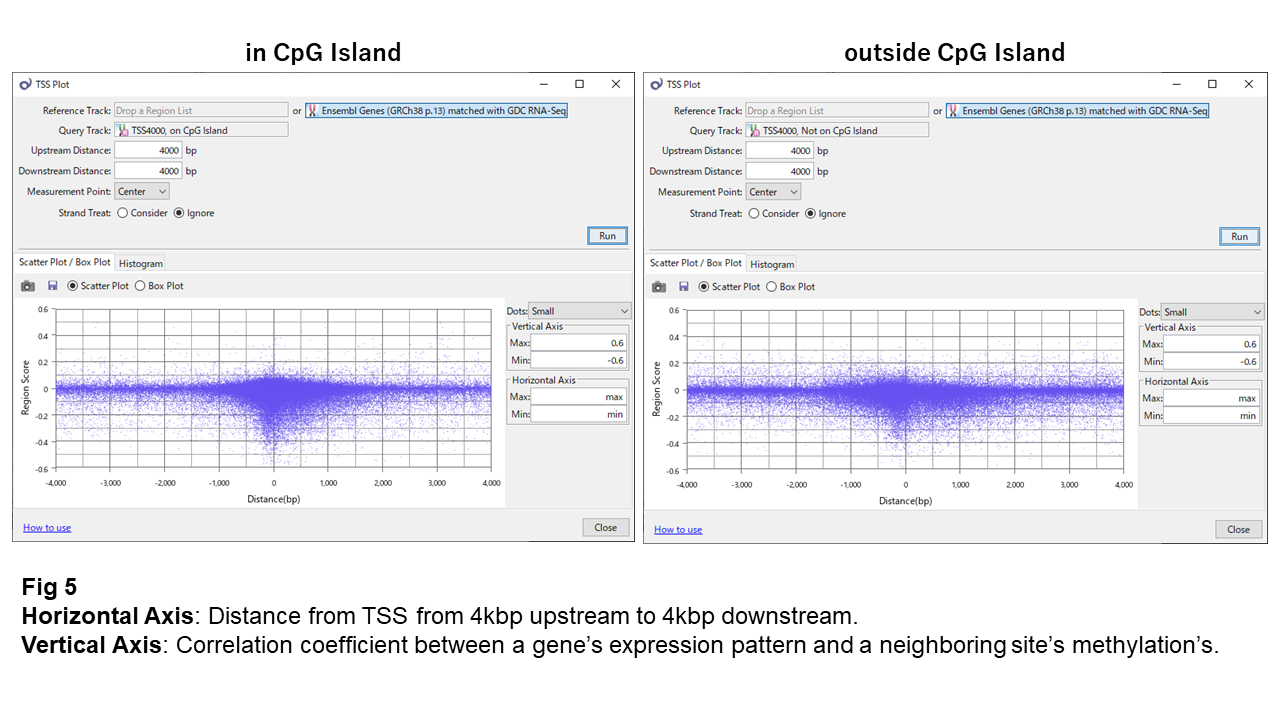
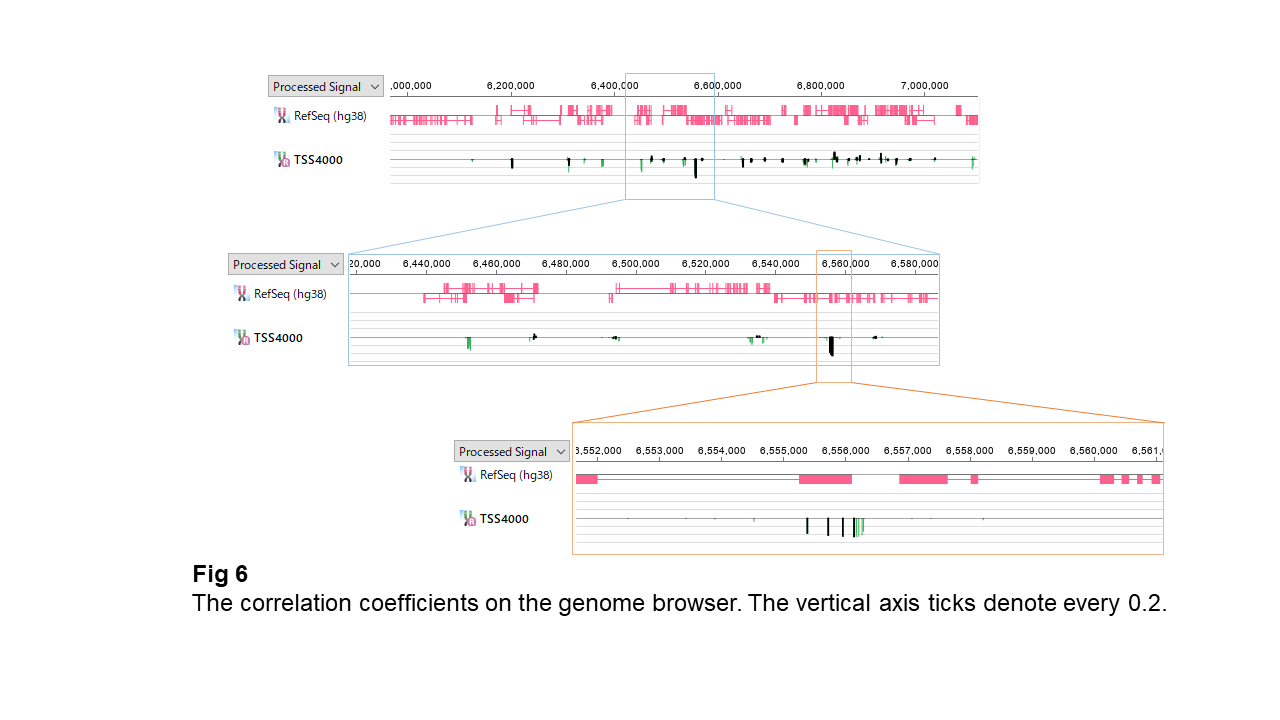
Fig. 6 shows these relationships along genomic coordinates. While most sites show correlations near zero, some exhibit moderate negative correlations.
These sites are not isolated—they often appear in small clusters, and adjacent sites tend to show similar correlation values.
This suggests that methylation sites very close to the TSS, including those in CpG islands and their immediate surroundings, tend to share similar relationships with gene expression.
CpG Island-Level Analysis
Given this local similarity, we next analyze methylation at the CpG island level.
We averaged beta values per CpG island and paired them with protein-coding genes whose promoters lie within 500 bp of the TSS. This yielded 11,035 gene–CpG island pairs.
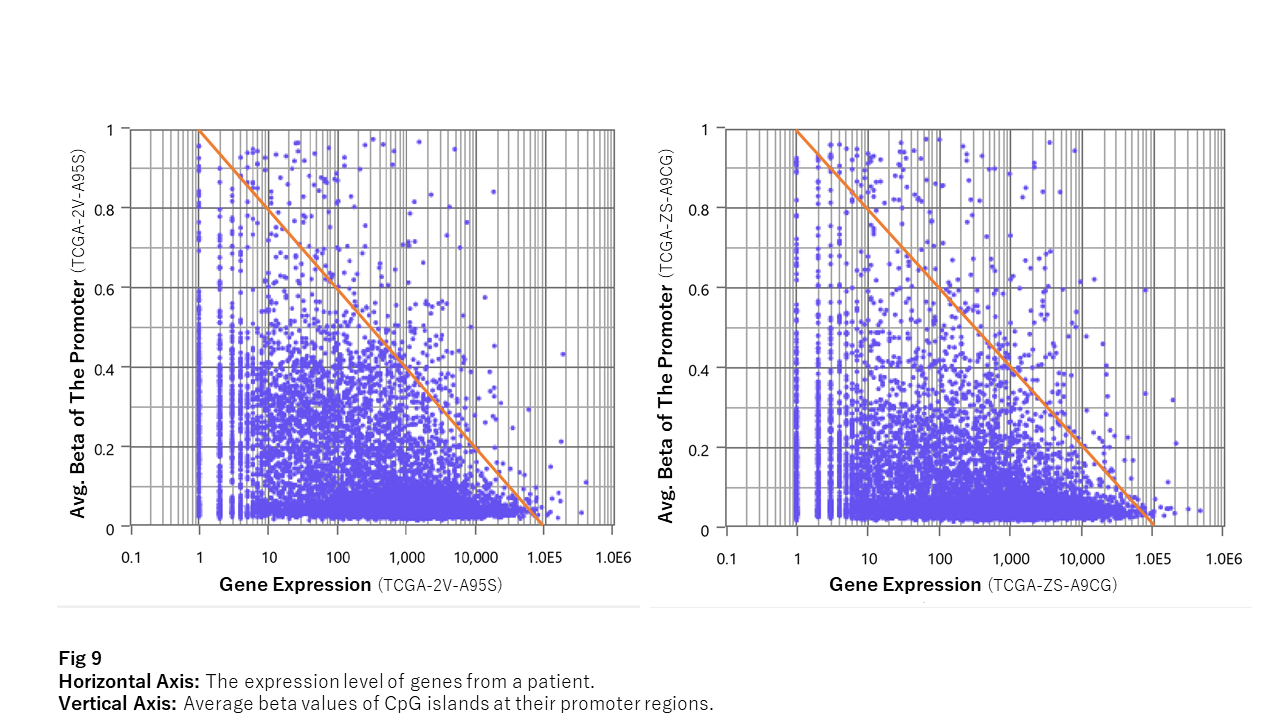
Fig. 9 shows the relationship between average beta values and gene expression.
The scatter plot forms a triangular distribution. What does this mean?
It reveals that while the well-known relationship—“methylation suppresses gene expression”—is indeed observed, the opposite—“demethylation enhances expression”—does not generally hold.
Global Patterns Across Patients
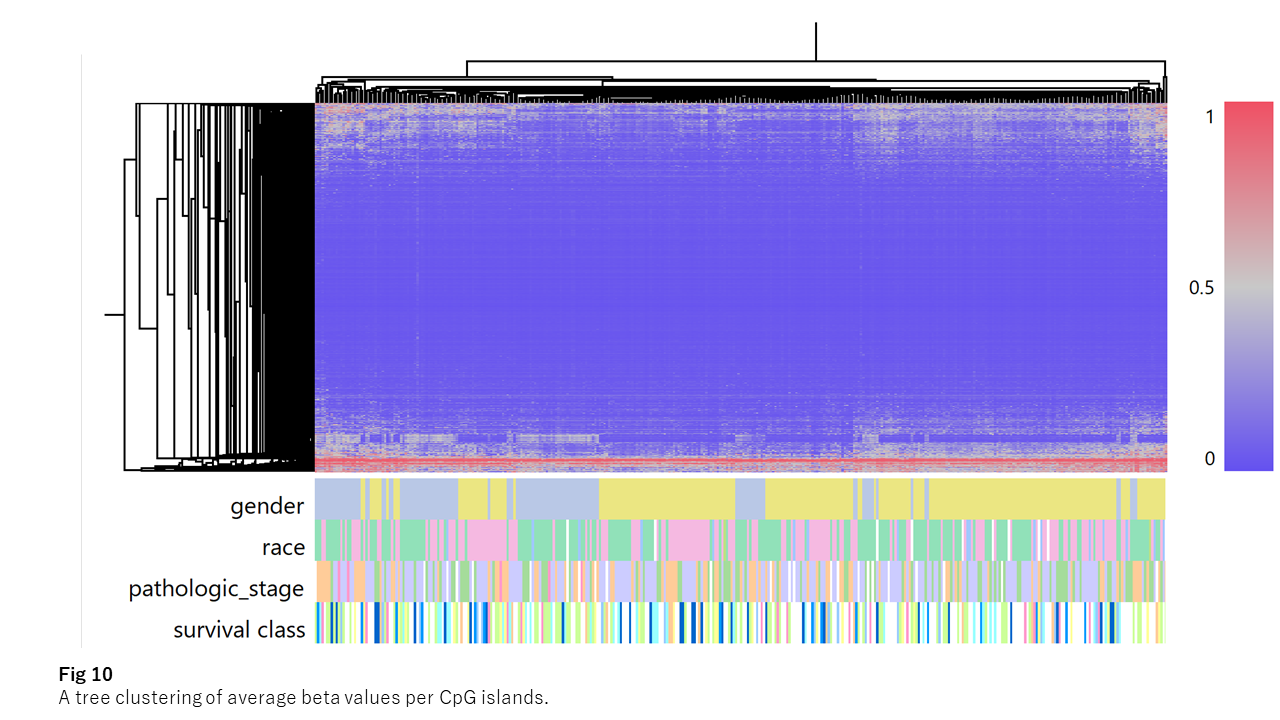
Fig. 10 shows average beta values across all patients.
Most CpG islands remain consistently hypomethylated across patients. A small subset remains highly methylated, possibly representing genes silenced during differentiation. While we do not explore this further here, examining these genes may provide additional insights.
The remaining ~4,000 CpG islands show variation across patients, suggesting potential involvement in gene regulation.
Correlation Depends on Variability
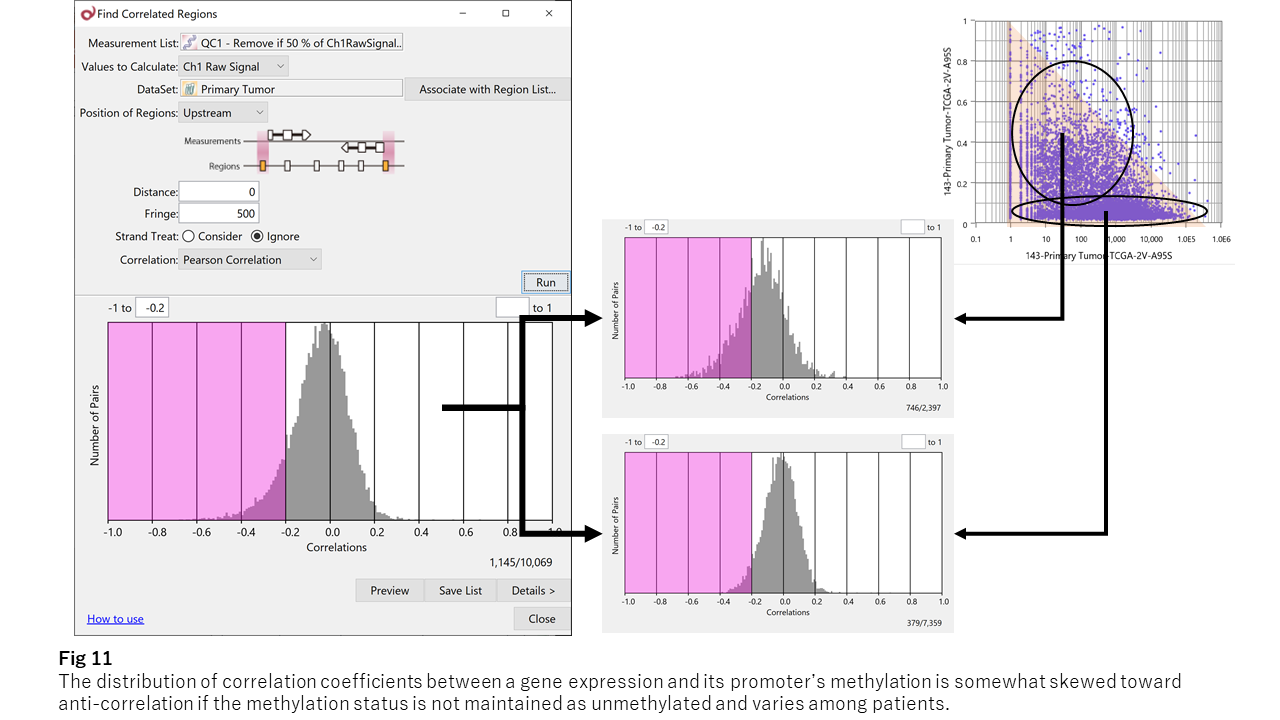
Fig. 11 shows correlations across CpG islands.
While centered near zero, the distribution is biased toward negative values, indicating a relationship between promoter methylation and gene expression.
When separating CpG islands into stable vs. variable groups:
- Stable (consistently unmethylated): no clear relationship
- Variable: weak inverse correlation becomes more apparent
Why Correlation Is Weak
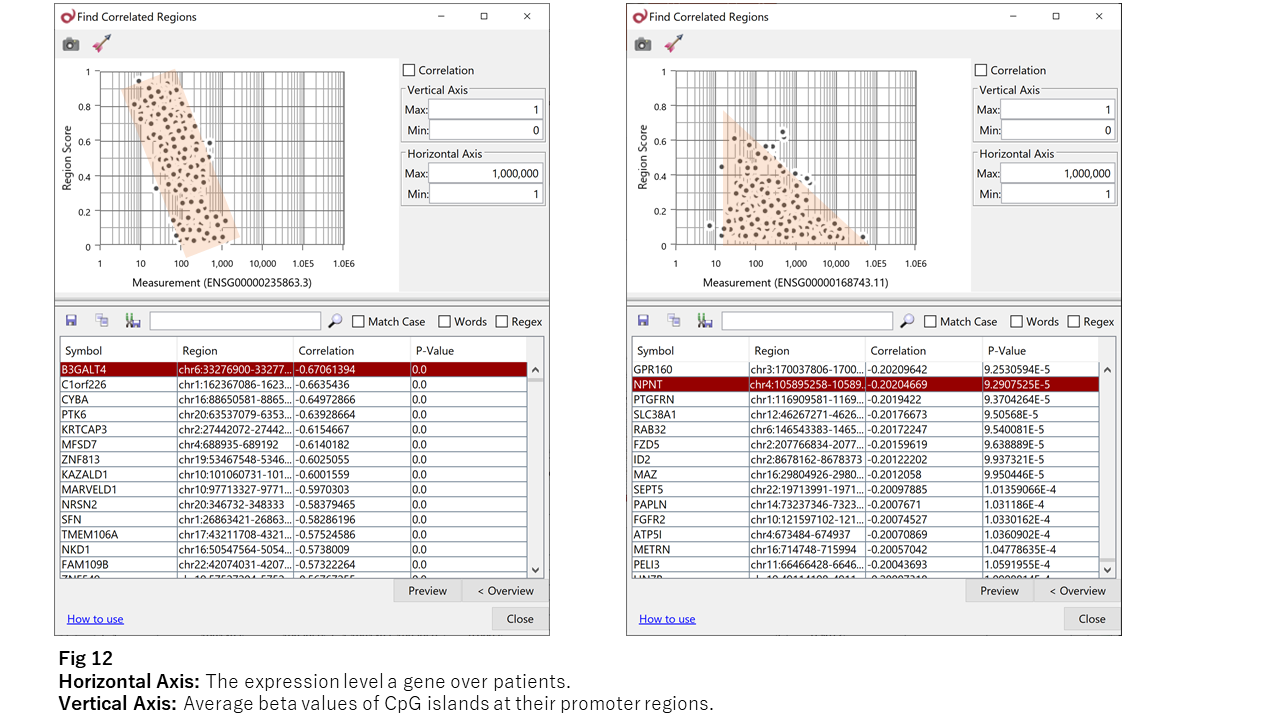
Strongly negatively correlated genes (e.g., B3GALT4; Fig. 12, left) are rare.
More commonly, genes show triangular distributions (e.g., NPNT; Fig. 12, right).
This suggests that:
- Increased methylation broadly suppresses expression
- Decreased methylation rarely leads to increased expression
This asymmetry likely explains why only weak negative correlations are observed overall.
As shown in Fig. 11, for many genes, promoter regions are biased toward a low-methylation state.
Within this range, gene expression varies widely, making any clear correlation difficult to observe.
However, when focusing on the subset of promoters where methylation levels vary, the relationship is not cancer-type specific.
Similar patterns are observed across multiple TCGA projects, and this has been consistently confirmed in our own analyses.
The key point is not to analyze all genes in bulk, but to focus on those with variable methylation and capture the resulting tendency toward weak inverse correlations.
Such relationships cannot be clearly captured without visualization.
We encourage you to explore whether the same patterns can be observed in your own data using scatter plots.
Spatial Specificity
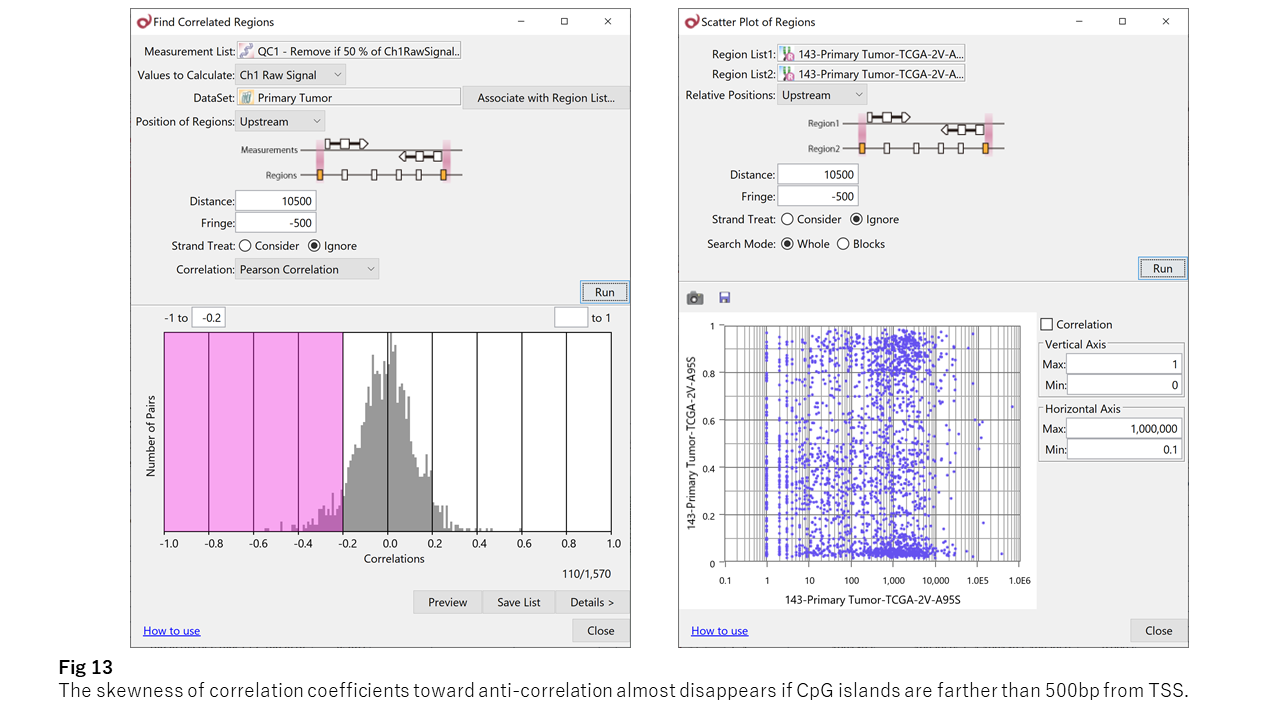
Importantly, this inverse correlation is specific to promoter regions within ~500 bp of the TSS.
Beyond this region, no clear bias toward inverse correlation is observed (Fig. 13).
However, individual sites may still show strong positive or negative correlations. Whether these are random or biologically meaningful remains an open question—one worth exploring directly in the data.
Conclusion: What Should We Read from Data?
As we have seen, the relationship between promoter methylation and gene expression is not simple.
While increased methylation is associated with suppression, decreased methylation does not necessarily lead to increased expression. The relationship is fundamentally asymmetric.
Today, AI can readily provide plausible explanations for such relationships. However, whether those explanations truly reflect real biological phenomena is another question.
That is why it is essential to examine data with your own eyes—and think about what it actually tells you.