What is the difference between miRNA-seq and RNA-seq?
The most important distinction is reproducibility.
This difference is most clearly reflected in how consistently results agree across datasets.
________________________________________
Technologies for measuring gene expression levels have reached a mature stage at least since around 2004. Therefore, data generated by skilled experimentalists can generally be considered reliable. In contrast, measuring microRNA (miRNA) expression remains technically challenging. It is important to recognize that miRNA expression data are not as reliable as gene expression data.
Here, we compare gene expression and miRNA expression data across multiple datasets of hepatocellular carcinoma (HCC).
________________________________________
Gene expression data show high reproducibility: consistent results across different technologies such as RNA-Seq and microarrays
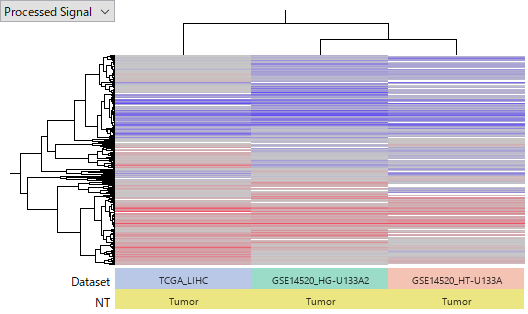
The heatmap below compares two datasets: TCGA-LIHC and GSE14520.
Gene expression data in TCGA were obtained by RNA-Seq, whereas GSE14520 was measured using two different microarray platforms (Affymetrix HG-U133A 2.0 Array and HT_HG-U133A Array), effectively treated here as three datasets.
All data were reprocessed from raw data and transformed into log2 ratios (Tumor vs Normal).
Red indicates upregulation in tumors, while blue indicates downregulation.
Despite being generated by different researchers and using different technologies, the patterns of up- and down-regulated genes are broadly consistent across all three datasets. This indicates that these data are highly reliable.
Such agreement across datasets is supported by the high reproducibility observed within each dataset.
________________________________________
Gene expression data are highly reproducible within datasets
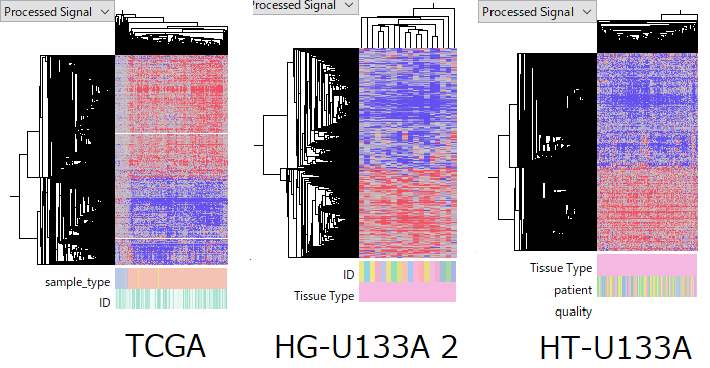
The TCGA-LIHC RNA-Seq dataset consists of 50 Normal and 370 Tumor samples.
The GSE14520 HG-U133A 2.0 dataset contains 18 paired Normal–Tumor samples, while the HT-U133A dataset includes 214 pairs.
Although a few questionable samples are present, these datasets are generally of high quality.
Within each dataset, tumor samples show broadly similar expression profiles. This supports the conclusion that these expression patterns reflect true biological states.
________________________________________
miRNA expression data show low reproducibility: inconsistent results across different measurement technologies
Now let us examine miRNA expression data.
Here, we compare five datasets generated using different technologies, including miRNA-Seq and microarrays.
In contrast to gene expression data, these results are strikingly inconsistent, with large discrepancies between datasets. Notably, even datasets generated using the same Affymetrix platform fail to produce consistent results.
This lack of agreement highlights the difficulty of generalizing conclusions from a single dataset. In other words, interpreting miRNA expression data requires far more caution than expected.
 ________________________________________
________________________________________
Are miRNA expression data consistent within individual datasets?
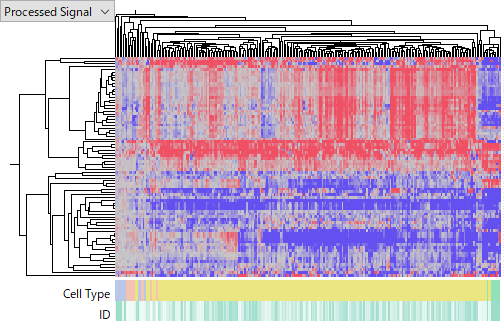
Among these datasets, the largest is TCGA-LIHC miRNA-Seq.
At first glance, the heatmap suggests that tumor samples show broadly consistent miRNA expression profiles. Looking only at this dataset, one might reasonably conclude that the experiment was successful and the results are reliable.
However, the very existence of such “apparently consistent” data highlights the fundamental difficulty of miRNA expression analysis.

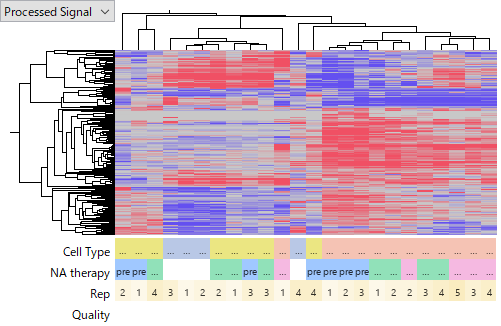
GSE110217 was measured using the Agilent Human miRNA v16 microarray.
Closer inspection reveals that signals in later replicates (samples 5–8) are clearly lower than in earlier ones (samples 1–4), suggesting a batch effect. These later samples were therefore excluded from the analysis.
This dataset includes three groups: Normal, non-HCC, and HCC.
Clustering results show that HCC samples form a distinct cluster, consistent with biological expectations. Thus, taken alone, this dataset appears valid.
However, once again, internal consistency within a dataset may itself highlight the difficulty of miRNA analysis.
The next two datasets, GSE74618 (Affymetrix miRNA v2 Array) and GSE115016 (Affymetrix miRNA v4 Array), were generated using different versions of Affymetrix platforms.
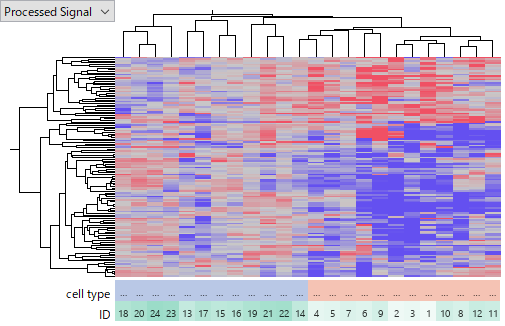
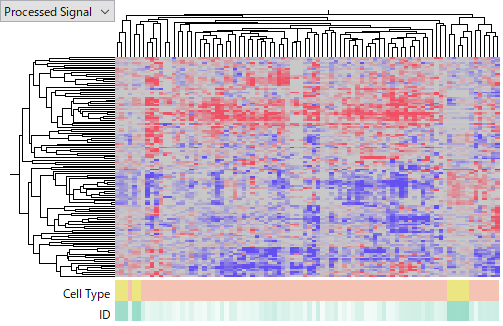
The next two datasets are GSE10694 (CapitalBio Mammalian miRNA Array) and GSE28854 (Milteny Biotec miRXplore miRNA Microarray), measured using different platforms.
GSE28854 contains only HCC patient samples and no control samples.
These datasets show greater variability than the previous ones, but internal consistency is not entirely lost.

________________________________________
As we have seen, a certain level of internal consistency exists within each dataset. However, even when such consistency is observed, it does not necessarily imply that the results are reliable.
Only by comparing multiple HCC datasets does it become clear how difficult it is to answer even a basic question:
Which miRNAs are truly upregulated or downregulated in HCC?
Comprehensive miRNA measurement is an evolving technology that still faces unresolved hurdles. When using miRNA datasets or relying on "up-regulated miRNA lists" from published papers, extreme caution is required.
________________________________________
Why do miRNA expression results fail to agree? Causes of low reproducibility
In contrast to RNA-Seq, miRNA-Seq results often vary significantly across datasets due to differences in experimental conditions and measurement systems.
This low reproducibility arises from several intrinsic properties of miRNAs.
Three major factors reducing reproducibility in miRNA expression data
Short sequences with high similarity
miRNAs are very short (~22 nucleotides), and differences between family members can be only a few bases.
This leads to cross-hybridization in microarrays and ambiguous mapping in sequencing, which can result in underestimation of true expression differences.
Library preparation bias
Differences in adapter ligation efficiency and RNA extraction kits are known to strongly influence miRNA quantification.
These biases help explain why results may appear consistent within a dataset but fail to agree across datasets.
Small number of miRNA species
The number of miRNAs is much smaller than that of genes (only a few thousand).
Therefore, normalization methods based on the assumption that “most genes do not change” are not always valid.
In situations where many miRNAs change simultaneously, it becomes difficult to define a stable baseline, making accurate comparison challenging.
As a result, standard normalization alone may be insufficient, and external controls such as spike-in RNAs are often required.
________________________________________
Conclusion
Overall, genome-wide measurement of miRNA expression remains a developing technology with significant unresolved challenges. When using miRNA datasets—or interpreting lists of differentially expressed miRNAs reported in the literature—careful consideration is essential.
Most importantly, the difficulty of miRNA expression analysis does not become apparent from a single dataset, but only emerges when multiple datasets are compared.
This is a problem that is easily overlooked.
Therefore, when planning miRNA experiments, more careful preparation—such as establishing a robust and consistent experimental system—and higher technical expertise are required compared to gene expression analyses.
Summary: Differences between RNA-Seq and miRNA-Seq
| Aspect | RNA-Seq (gene expression) | miRNA-Seq |
|---|---|---|
| Reproducibility | High | Variable and condition-dependent |
| Cross-dataset consistency | Generally consistent | Often inconsistent |
| Measurement stability | Relatively stable | Strongly affected by technical bias |
| Assumptions for analysis | Generally valid | Often difficult to satisfy |
Download the data for your Subio Platform
If you want to look closer these data sets with Subio Platform by yourself, download the SOA file which works like a bundle of SSA files.
Open "Import Archive..." under Platform menu, and select the SOA file. Subio Platform automatically shuts down when it completes importing. Please restart the software to see the all data sets.
________________________________________
[2026 Update] Never Ask AI, "Which miRNAs are Up-regulated in X?"
While AI-driven analysis is becoming more common in 2026, it is dangerous to blindly accept the "answers" provided by AI when the quality of the underlying data varies so drastically.
For instance, if you ask an AI, "Which miRNAs are up-regulated in HCC (Hepatocellular Carcinoma)?", it will confidently present a list extracted from past publications. However, the technical reality demonstrated in this article is that results vary significantly from paper to paper, often incorporating data of questionable reliability.
Furthermore, AI models can extract patterns even from noisy data, sometimes identifying features that may not reflect true biological signals. Ultimately, the responsibility for determining whether these results are biologically valid lies not with the AI, but with the analyst. ________________________________________
It is also important to note that the same caution applies to databases such as miRmine and miRNAMap, which aggregate expression levels across various tissues. While these databases can be useful, they often integrate data generated under different experimental conditions, making it difficult to distinguish biological differences from technical variation.
For omics data analysis, it is essential to first examine the data directly, rather than relying solely on AI-generated answers.