This page explains the limitations and data quality issues of single-cell RNA-Seq (scRNA-Seq) analysis in an accessible way.
Introduction: Have You Ever Looked Behind the “Beautiful Plots”?
Single-cell RNA-Seq (scRNA-Seq) has now become a standard approach in omics analysis. Colorful plots generated by tools such as Loupe Browser can look as if they reveal the true state of cells, almost like images from immunostaining or FISH.
However, we also need to look behind that visual appeal and consider the instability of the underlying data. When the data are examined in detail using Subio Platform, we can see that there is an underlying “fragility” that cannot be fully corrected computationally, along with “chance” signals that may have been detected only by accident.
1. The Problem of Sequencing Depth in Single-Cell RNA-Seq and the Illusion of “Cellular Identity”
This is the warning we raised in 2019, when single-cell RNA-Seq was beginning to spread widely.
In protocols such as 10X Genomics, the number of reads per cell is only on the order of tens of thousands. Compared with conventional bulk RNA-Seq, the difference is overwhelming.
A small number of reads means that the dynamic range—the measurable range—is extremely narrow. If you display read count histograms for each sample in Subio Platform, it becomes immediately clear that there are large differences in sequencing depth, that is, how many read counts were obtained for each cell.
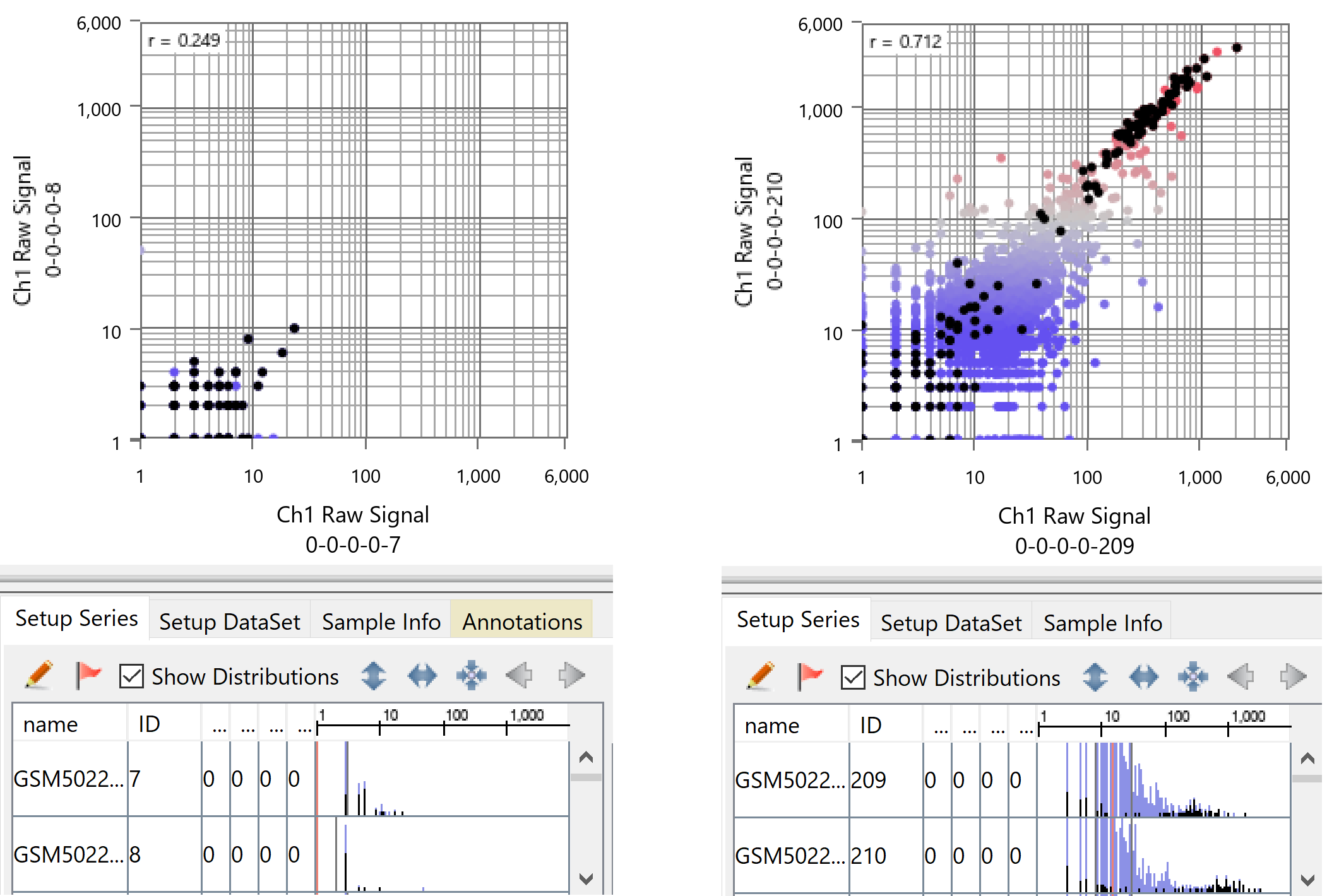
The figure above shows scatter plots comparing cells with the shallowest depth and cells with the deepest depth, created from GSE164898. Black dots indicate ribosomal protein genes, which are expected to be highly expressed in all cells. This figure shows the following:
- Cells with shallow depth: Even ribosomal protein genes are barely detected, and only some of them are captured.
- Cells with deep depth: At most, a few additional genes with expression levels comparable to ribosomal protein genes are measured.
What this tells us is that the technology itself carries a structural risk: simple “differences in sequencing depth” may be interpreted as “cellular identity.”
There is another fundamental principle that must not be forgotten: “Not detected” does not mean “not expressed.” It may simply mean that the transcript was not detected by chance.
The “beautiful plots” generated by analysis tools are actually built from accumulations of highly unstable data. We need to understand this risk.
2. Do Not Ask AI Only for “Solutions” — A 2026 Perspective
Now let us look at this issue from the perspective of 2026. The decisive difference from 2019 is that anyone can now consult AI about data analysis.
If you ask AI, “What are the limitations of single-cell RNA-Seq?”, it will likely return a reasonable answer. But here is something you should try next. Ask AI: “How can I correct batch effects in single-cell RNA-Seq?” AI will immediately suggest sophisticated solution methods such as MNN or Harmony.
If you do not first understand the “quality limitations” of the data, you may accept that answer as it is. But once you understand the fundamental “fragility” of the data, a question should arise:
“Can data with missing signals really be ‘corrected’ by computation alone?”
Once this question emerges, you should be able to ask AI more specific and critical questions about the characteristics and limitations of each method. Instead of asking only for solutions — the “How” — we need to compare those solutions against the underlying structure of the problem — the “What.” Noticing this contradiction and having a healthy sense of doubt — “Is this really true?” — is the critical thinking required for analysts to survive in the AI era.
The Mirage Reflected by Advanced Normalization
Between 2019 and 2026, there have also been advances in correction methods for scRNA-Seq. CPM (Counts Per Million) is commonly used to compensate for differences in depth. However, in cells with extremely few reads, the accidental detection of a single read can have a large effect and amplify variation in expression values. This “amplification of noise in low-input data” is not a problem unique to single-cell data. As shown in Case Study 403, it is a universal pitfall in data analysis that also appears in low-expression gene analysis in bulk RNA-Seq.
Methods such as TMM and median-of-ratios are now often recommended to address this issue, although they were originally designed for bulk RNA-Seq. However, when applied to scRNA-Seq data, they cannot necessarily be regarded as safe solutions. This is because the “corrected data” produced by sophisticated algorithms are not necessarily closer to the truth than simple CPM, and evaluating their validity is not easy. These methods depend on the statistical assumption that “most genes are not changing.” But remember: in scRNA-Seq, only a very limited set of extremely highly expressed genes can be detected stably. If we can observe only this small subset of genes, the validity of that assumption becomes unstable.
The practical attitude is not to blindly trust a particular method, but to compare the results of multiple normalization methods side by side. Precisely because we do not know which result is correct, we need to compare multiple “corrected datasets” with the raw count data, and ask under which conditions the data distortion appears smallest, or the observations can be explained most convincingly. In the end, we may have no choice but to judge it with our own eyes.
The question of how to interpret correction results when the shapes of data distributions differ substantially is also important in bulk RNA-Seq. As a case study applying multiple normalization and correction methods to bulk RNA-Seq data, please see Limitations of Batch Effect Correction and Normalization in RNA-Seq|Comparing ComBat, VST, TMM, and Quantile Normalization .
Incidentally, if you want to view raw scRNA-Seq data in Subio Platform, download the matrix file from GEO and ask ChatGPT: “Create a Python script that outputs (file name) as a dense matrix TSV, with genes as rows and cells as columns.” If you run the generated script in Jupyter, you can create a file that can be imported directly into Subio Platform. Thanks to AI, this kind of “work” has truly become easy. For details, please also refer to the related article “Generate Code with ChatGPT and Run It in Jupyter” .
Even in 2026, there are still points that require caution regarding the measurement sensitivity and stability of scRNA-Seq data, as well as the interpretation of corrected results. We should not assume that everything is fine simply because we are using the latest advanced methods. While expecting future improvements not only in correction algorithms but also in the sensitivity and stability of measurement systems, it is important to maintain a critical-thinking mindset.
scRNA-Seq Results Should Be Treated Not as Definitive Facts, but as Structural Hypotheses
The important point here is not to deny the value of single-cell RNA-Seq. Single-cell RNA-Seq is a very powerful method for exploring cellular heterogeneity and identifying candidate unknown cell states or rare populations.
However, scRNA-Seq data are interpreted only after passing through many preprocessing steps and assumptions, including dropout, low capture efficiency, differences in total read counts and detected genes among cells, normalization, batch correction, dimension reduction, clustering resolution, and cell type annotation.
For this reason, UMAP separation, trajectories, cell states, latent spaces, and cluster structures after data integration should not be understood as facts directly observed in the data itself. Rather, they should be understood as interpretation candidates obtained under specific preprocessing conditions and analysis parameters.
In particular, batch correction and data integration are useful methods for reducing unwanted technical differences. At the same time, they may also weaken biologically meaningful differences or bring together cell populations that should originally be considered separately. In other words, the corrected space is not “the truth itself,” but also a data space reconstructed by the correction algorithm and its assumptions.
Therefore, it is scientifically sound to treat scRNA-Seq results not as “definitively discovered structures,” but as structural hypotheses supported under specific conditions. What matters is whether those structures remain stable when analysis conditions are changed, whether they can be confirmed using independent data or other methods, and whether they are consistent with known marker genes and biological knowledge.
This does not mean that scRNA-Seq is meaningless. On the contrary, scRNA-Seq is extremely powerful for hypothesis generation. However, to claim its results as biological facts, we need to evaluate them carefully, not only based on visualization or clustering results, but also by considering reproducibility under multiple conditions, independent validation, and experimental confirmation.
What matters is neither blindly trusting scRNA-Seq nor rejecting it entirely. We should use it while understanding its limitations and assumptions, and while remaining aware of where observation ends and interpretation begins.
Related Topics
- A Practical Guide to RNA-Seq Analysis Without Black Boxes — Becoming an analyst who can think and make decisions in the AI era
- RNA-Seq Data Quality Depends on RNA Amount
(With very small amounts of RNA, it is difficult to obtain data quality comparable to ordinary bulk RNA-Seq.)