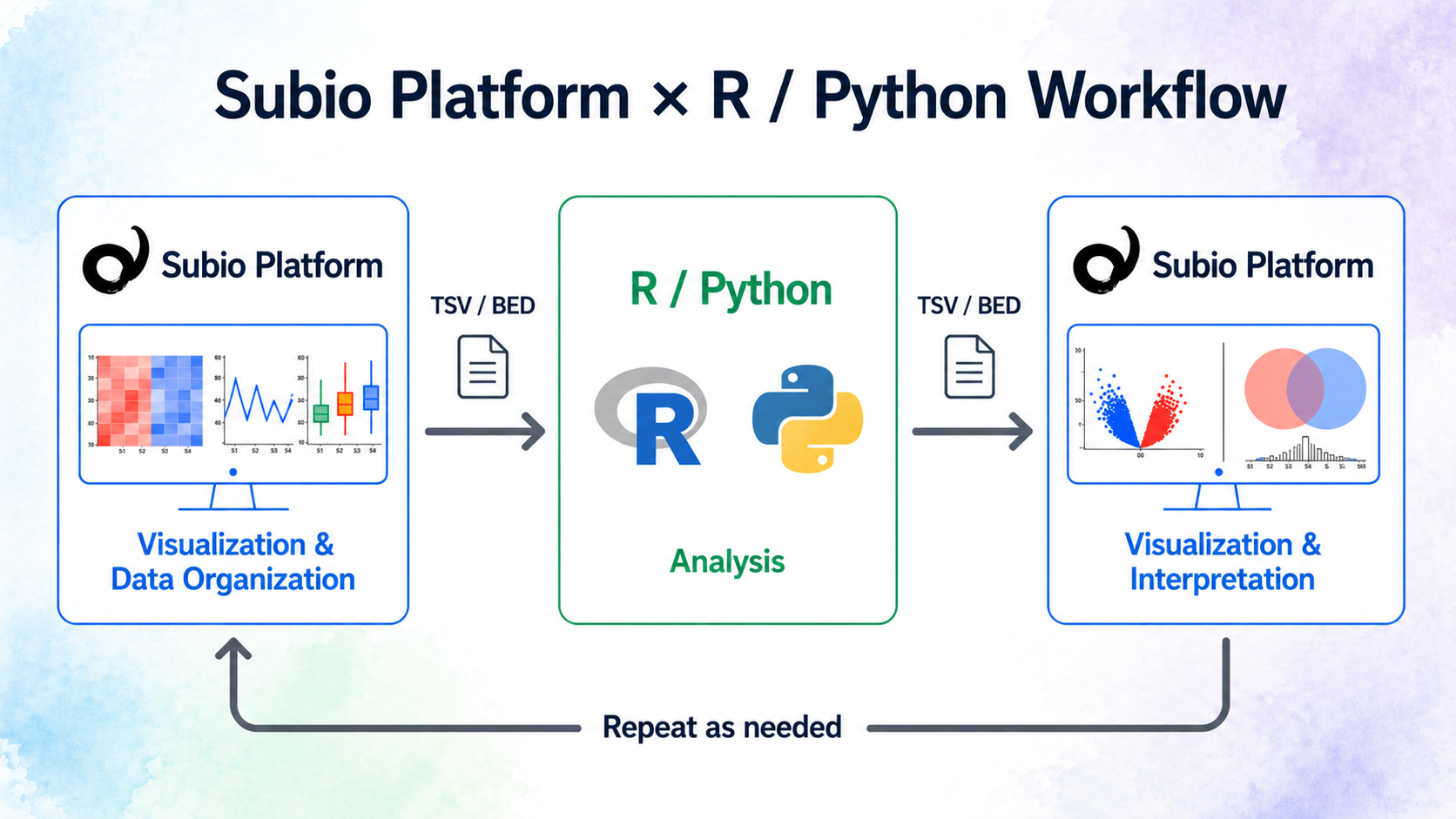
Subio Platform allows you to intuitively visualize, organize, and preprocess your data.
On the other hand, R and Python are well suited for advanced statistical analysis and custom workflows.
By combining them, you can build a workflow that helps you analyze while understanding your data.
This page explains the data formats used to connect Subio Platform with R / Python, and how this workflow can be used in practice.
Why Combine Subio with R / Python?
Subio Platform is strong in data visualization, data organization, and data management.
R and Python are strong in flexible statistical analysis and custom computation.
By combining them, you can:
- Check the state of your data through visualization
- Run flexible statistical analyses
- Bring the results back for visualization and interpretation
- Store analysis results and reuse them for further exploration
This makes it possible to choose analysis methods flexibly while turning your results into reusable research assets.
Data That Can Be Exported from Subio Platform
Subio Platform can export the following types of data for use in R / Python.
| Data | Format | Main Uses |
|---|---|---|
| Gene Count / TPM / FPKM (raw or preprocessed data) |
TSV | Differential expression analysis (DESeq2 / edgeR), PCA, UMAP, clustering, machine-learning-based classification and prediction |
| Sample information (groups, conditions, etc.) |
TSV | Group settings for differential analysis, classification and labeling (including supervised machine learning), batch information management |
| Annotated gene lists | TSV | Enrichment analysis, network analysis |
| Genomic region lists with values (ChIP-Seq, methylation, CNV, etc.) |
BED | Genome-position-based analysis, multi-omics integrative analysis |
These data can be read directly in R or Python.
Data That Can Be Imported into Subio Platform
Results generated in R or Python can be imported into Subio Platform for visualization and interpretation.
For example, the following types of data can be imported and used for visualization and interpretation:
-
Results of statistical differential expression analysis (P-values, FDR, etc.)
By importing them as a Measurement List, you can visualize differentially expressed genes and compare or combine conditions using Venn diagrams. -
Normalized or corrected data
By importing them as separate samples, you can visually compare the original data with corrected data, or compare different correction methods. -
Classification and characterization results from clustering or machine learning
These can be imported as sample attributes and used for visualization, comparison, validation, survival curve analysis, and related workflows. -
Genomic regions with associated numerical values
These can be visualized in the genome browser as bar graphs or heatmaps, and used to examine relationships or correlations with gene expression. -
PCA loadings (coefficients for each gene), or component vectors obtained from methods such as NMF or ICA
These can be used as Profiles for visualization in Scatter Plot of Samples.
Note: In linear methods such as PCA and NMF, these coefficients can be used to calculate sample scores or positions.
Note: In nonlinear methods such as UMAP and t-SNE, equivalent coefficients are not defined, but the resulting sample coordinates can still be used for visualization.
By importing these data, you can visualize and interpret analysis results from multiple perspectives.
Common Points to Note
When connecting Subio Platform with R or Python, keep the following points in mind:
- Subio Platform supports TSV format, but not CSV format.
- Gene IDs and sample names can be handled flexibly, but it is recommended to use consistent IDs and naming rules to keep the relationships clear during analysis and visualization.
If these relationships are not handled properly, sample or gene mappings may become misaligned, which can lead to incorrect interpretation of the results.
Using AI Tools Such as ChatGPT
In recent years, it has become increasingly common to use ChatGPT to generate R or Python code and run analyses.
By organizing your data in Subio Platform and generating code only for the necessary parts, you can:
- Minimize the amount of code you need
- Make error correction easier
- Reduce maintenance effort
Building a large pipeline with AI all at once can be risky. However, using small programs in combination is a realistic and efficient approach.
Next Step
For a complete guide to RNA-Seq data analysis, please see the tutorial below:
→ RNA-Seq Data Analysis Tutorial
You can follow the tutorial step by step using real data while combining Subio Platform with R / Python.