Introduction
You can easily import the TCGA methylation and RNA-Seq data sets with Subio Platform. (Watch tutorials " how to import RNA-Seq data " or " DNA methylation data " for import operations.) This case study is about practical data analysis as integrating DNA methylation and gene expression data.
I chose TCGA-LIHC (Liver Hepatocellular Carcinoma) dataset for an example. Let's see the summary of DNA methylation data. I extracted probes located within 4kbp from TSS of genes measured in RNA-Seq to examine.
Please be noticed that they measured DNA methylation with Illumina's Infinium HumanMethylation450 BeadChip, which they intensively selected probes near TSS. Consequently, the following TSS plots have a bias in density around TSS, but it may not reflect the biological fact.
The summary of TCGA-LIHC methylation data.
Fig 1 shows the distribution of beta values against the TSS's relative distance. The beta value indexes the degree of methylation from 0 (unmethylated) to 1 (methylated). The two plots represent sites inside and outside CpG islands, respectively.
Most sites inside CpG islands look kept unmethylated though, some near the TSS can be somewhat methylated. On the contrary, those outside CpG islands look mostly methylated, except regions around TSS (within 1 ~ 1.5kbp) where they are primarily unmethylated.
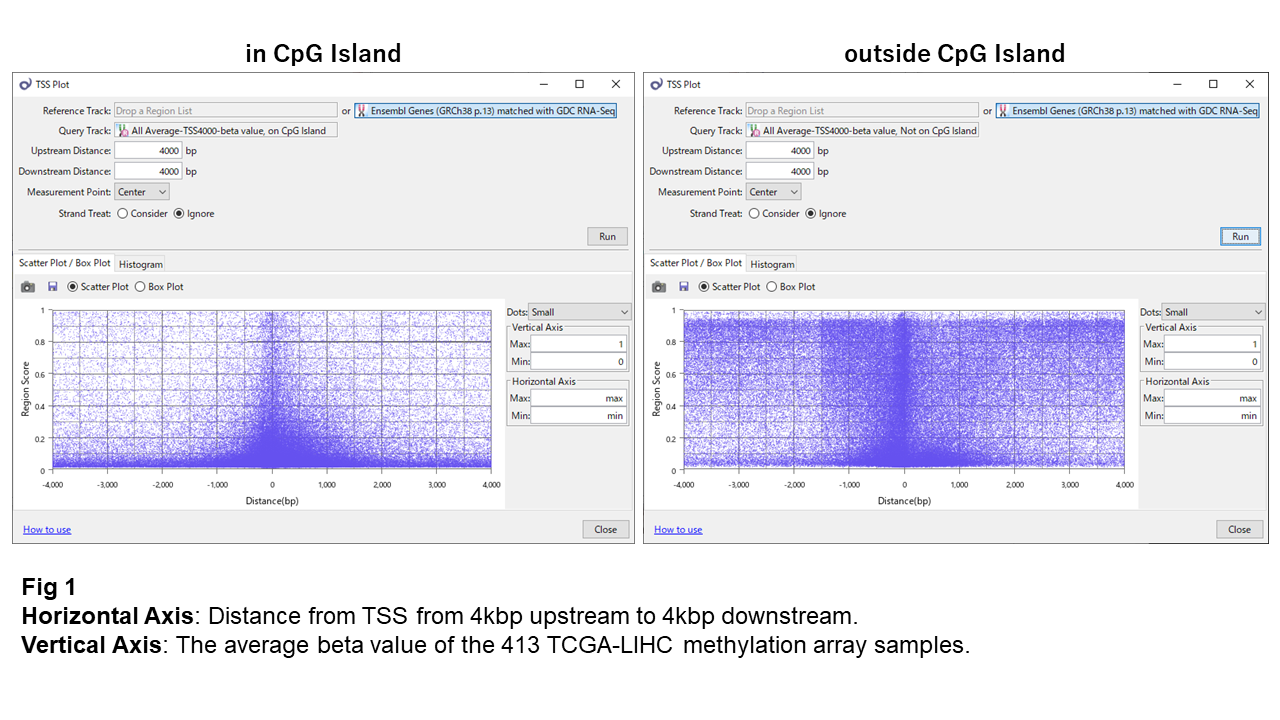
Fig 2 shows the differences between Solid Normal Tissue and Primary Tumor against the TSS's relative distance. The positive value on the vertical axis indicates hyper-methylation in Tumor, and the negative does hypo-methylation.
Sites in CpG islands look strictly force kept the methylation levels in general, except the area near TSS. The change there is mostly toward hyper-methylation. On the other hand, those outside CpG islands also maintain the methylation levels, but maybe less strictly. And the direction is primarily toward hypo-methylation.
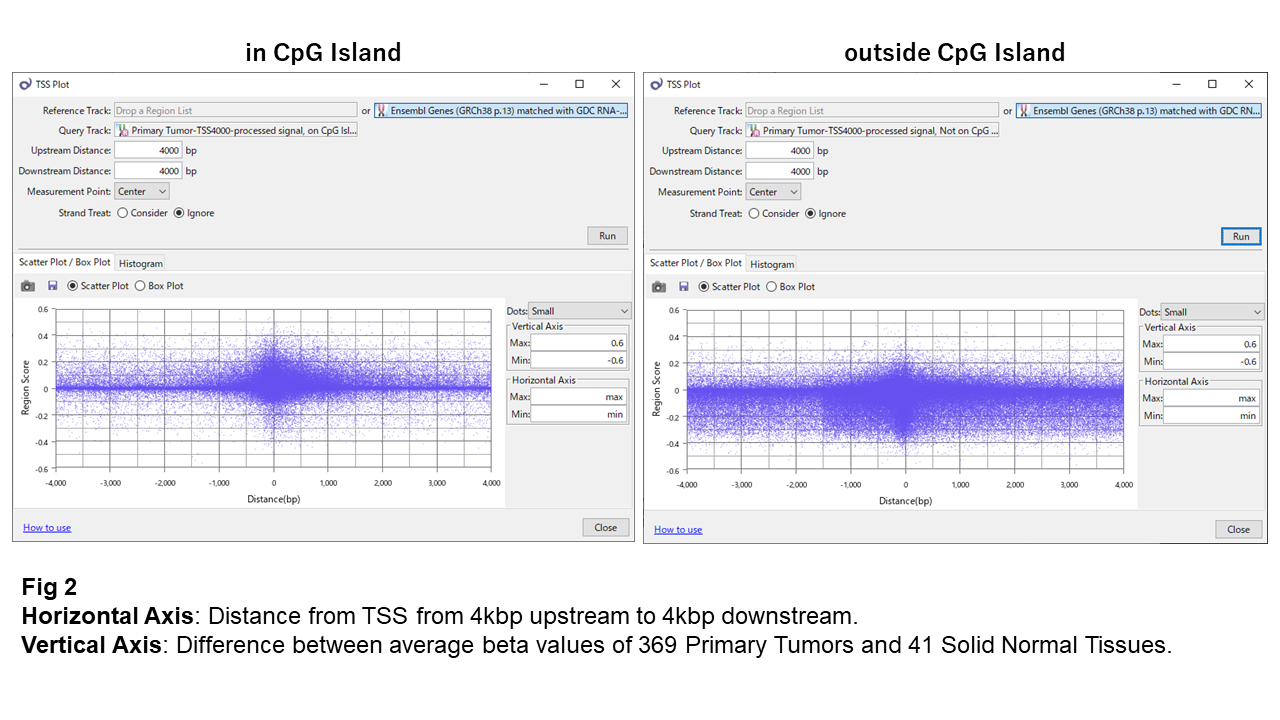
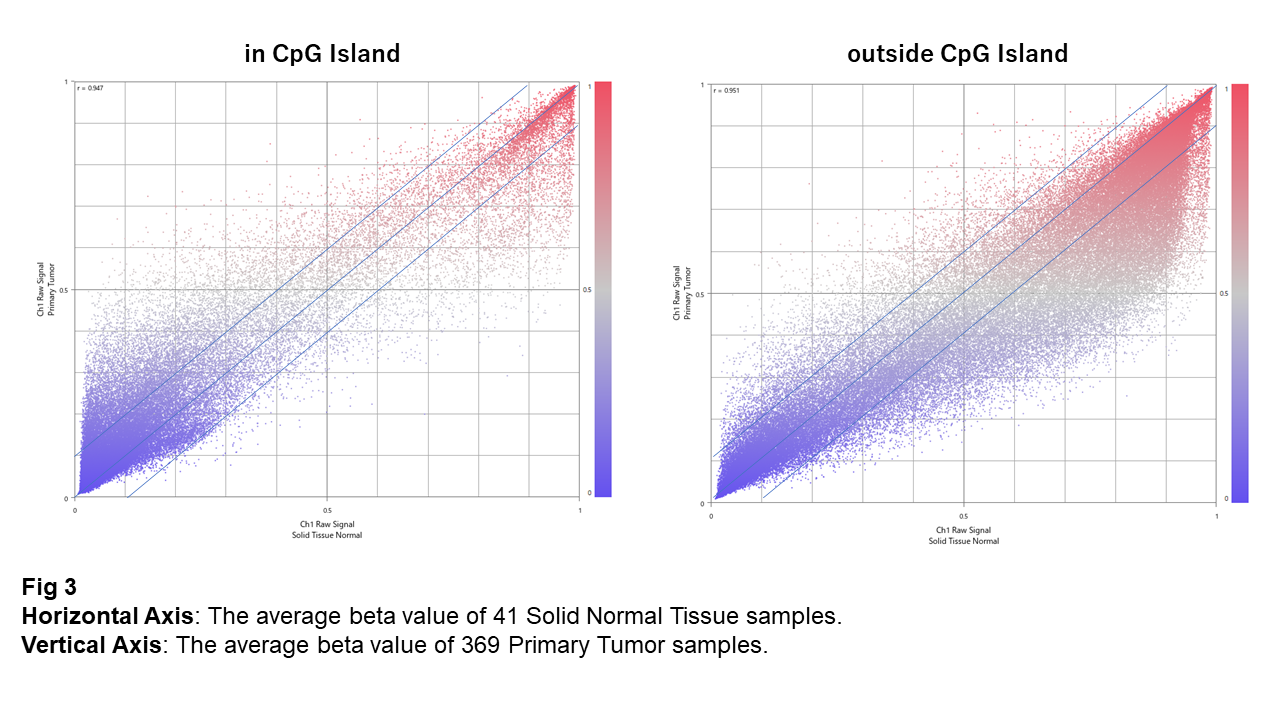
Fig 3 compares the average beta values between Normal and Tumor. In CpG islands, the remarkable change is the hyper-methylation of unmethylated sites. On the contrary, the dominant difference for sites outside CpG islands is the hypo-methylation of highly-methylated ones.
The correlation between the methylation and expression patterns.
And here's the thing. Let's see how much the DNA methylation relates to the neighboring gene's expression levels.
Fig 4 represents the correlation coefficients in pairs of a site's methylation pattern and a neighboring gene's expression pattern. The coefficients strongly converge to zero, indicating that almost all couples show no-correlation. But still, you see the distribution is left-skewed. Some pairs seem to have weak anti-correlation.
I calculated the same as changing the order of samples (Negative Control) to confirm that anti-correlation skewness disappears. Consequently, the anti-correlations are significant even though they are weak. I extracted anti-correlated pairs with a threshold of -0.3.
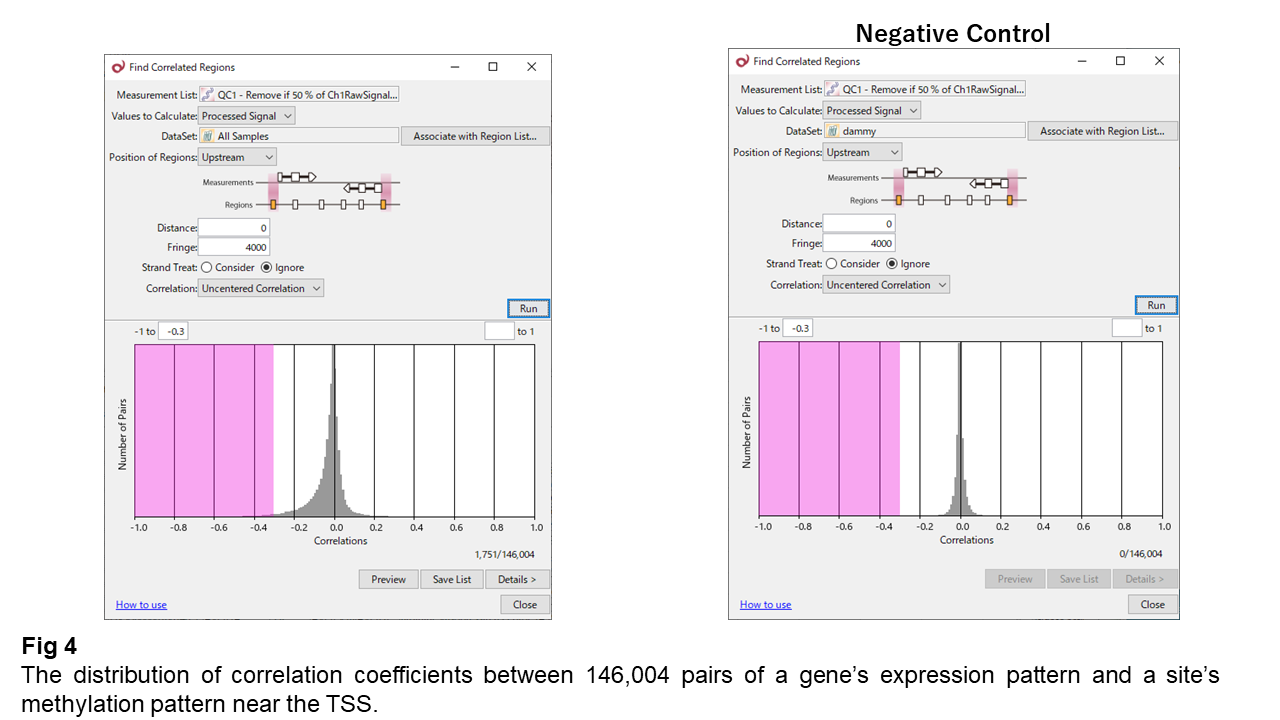
Fig 5 displays the correlation coefficients against the TSS's relative distance. It suggests the anti-correlated sites are near TSS (within 300bp) intensively and are sparse as they go distant. The number of anti-correlated sites in CpG islands is 1,007, and it is 715 for outside. Because they designed more probes for CpG islands in the array, I calculated the percentage of going less than -0.3. They were 1.4% and 1.3% respectively, and they are not so different.
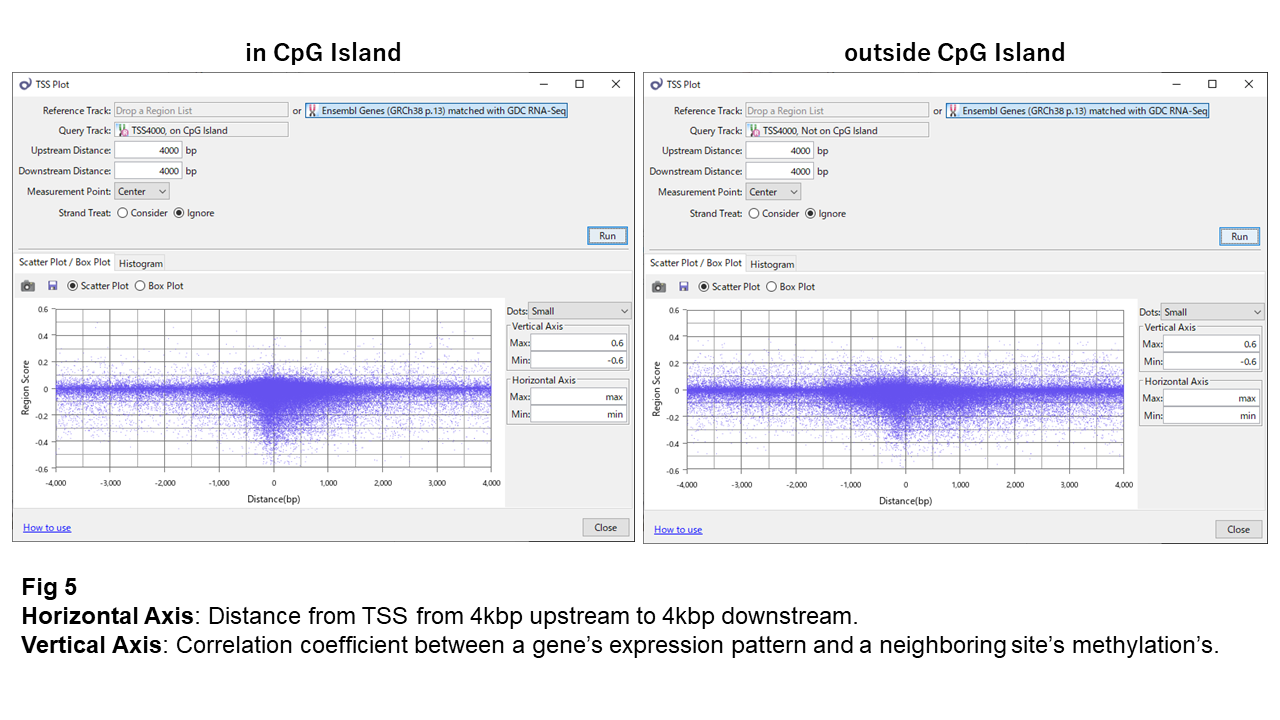
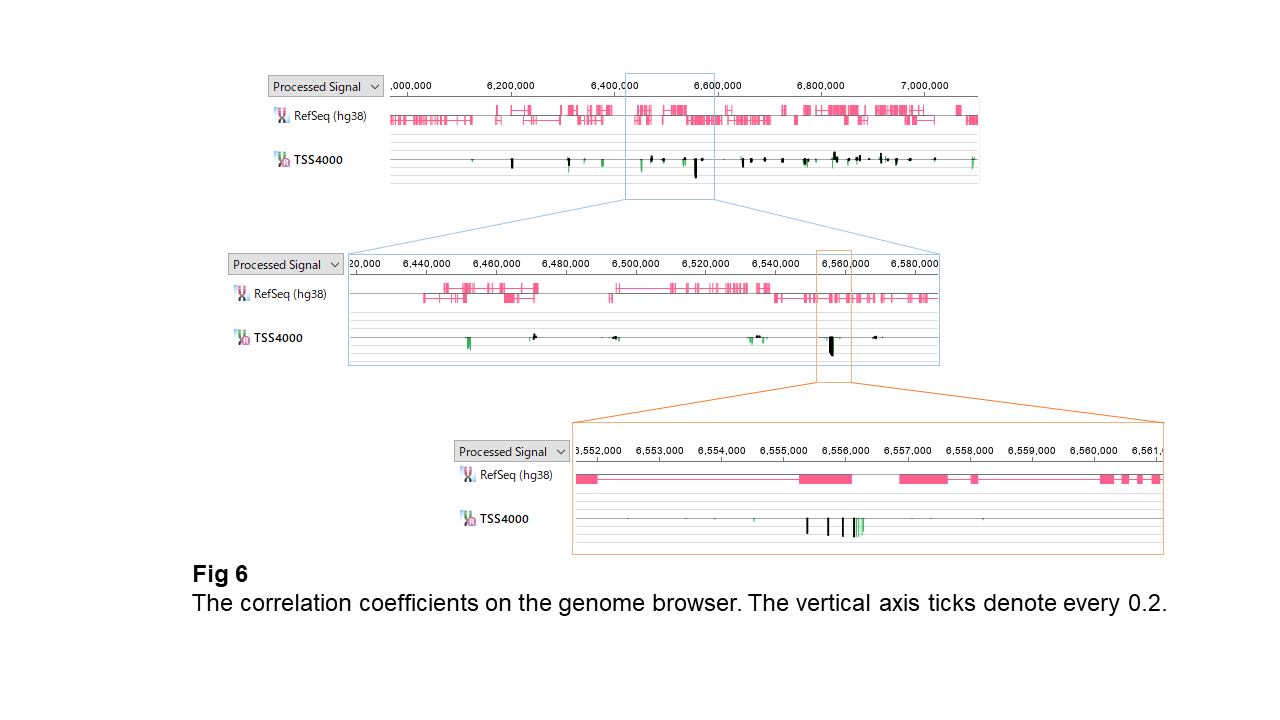
Fig 6 displays genes and methylation sites according to the genomic coordinate. The bar chart at each site represents the correlation coefficients between the methylation pattern and the neighboring gene's expression pattern. Black bars are for ones in CpG island, and green for outside. Though most bars are around zero, a few reach under the -0.2 line. They can be either in or outside CpG islands. Remarkably, the closely-neighboring sites share almost the same coefficient values.
The effect of the methylation status of CpG island on the neighboring gene's expression level.
We examined the individual methylation sites. And now, let's see the methylation status per CpG island. First, I calculated the average beta values per CpG island and coupled them with the gene whose TSS locates within 500bp. And then, I extracted 11,035 pairs of a CpG islands neighboring protein-coding genes measured by the TCGA-LIHC RNA-Seq.
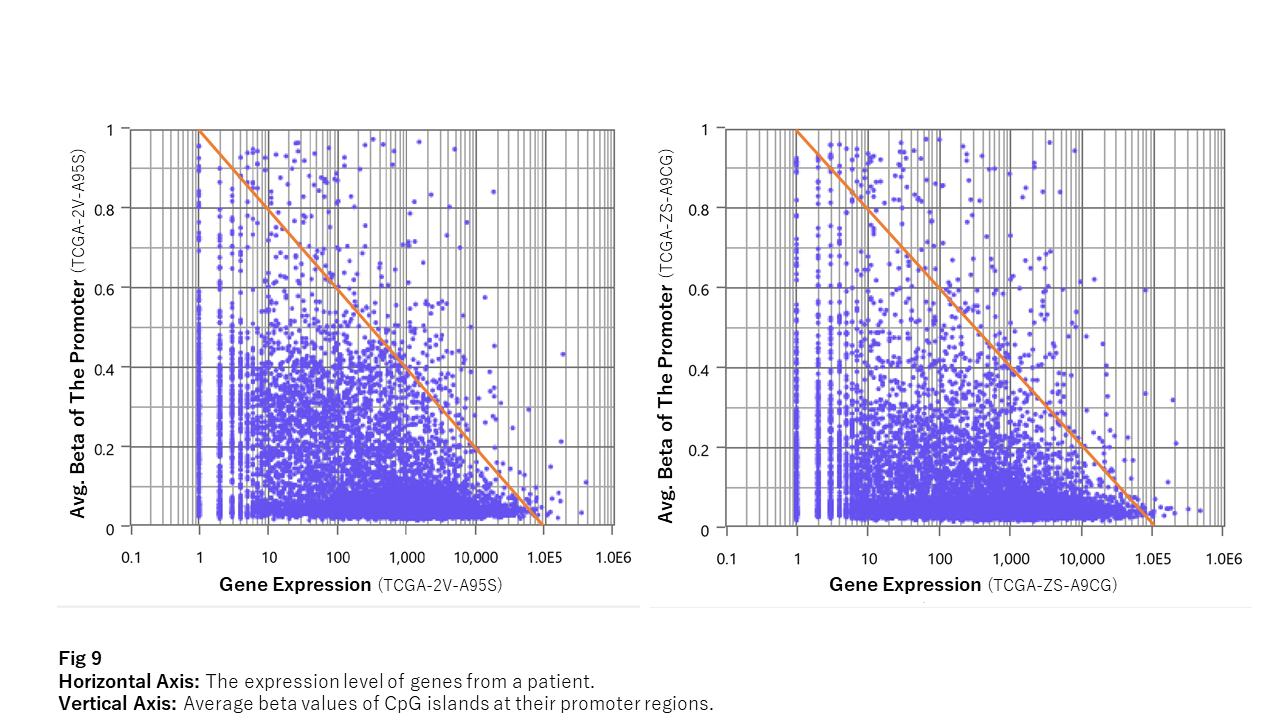
Fig 9 shows the relationship between the average beta values of CpG islands and the expression level of the neighboring gene per patient (TCGA-2V-A95S and TCGA-ZS-A9CG). The triangle shape of plots implies that methylation at the promoter limits the adjacent gene’s expression level.
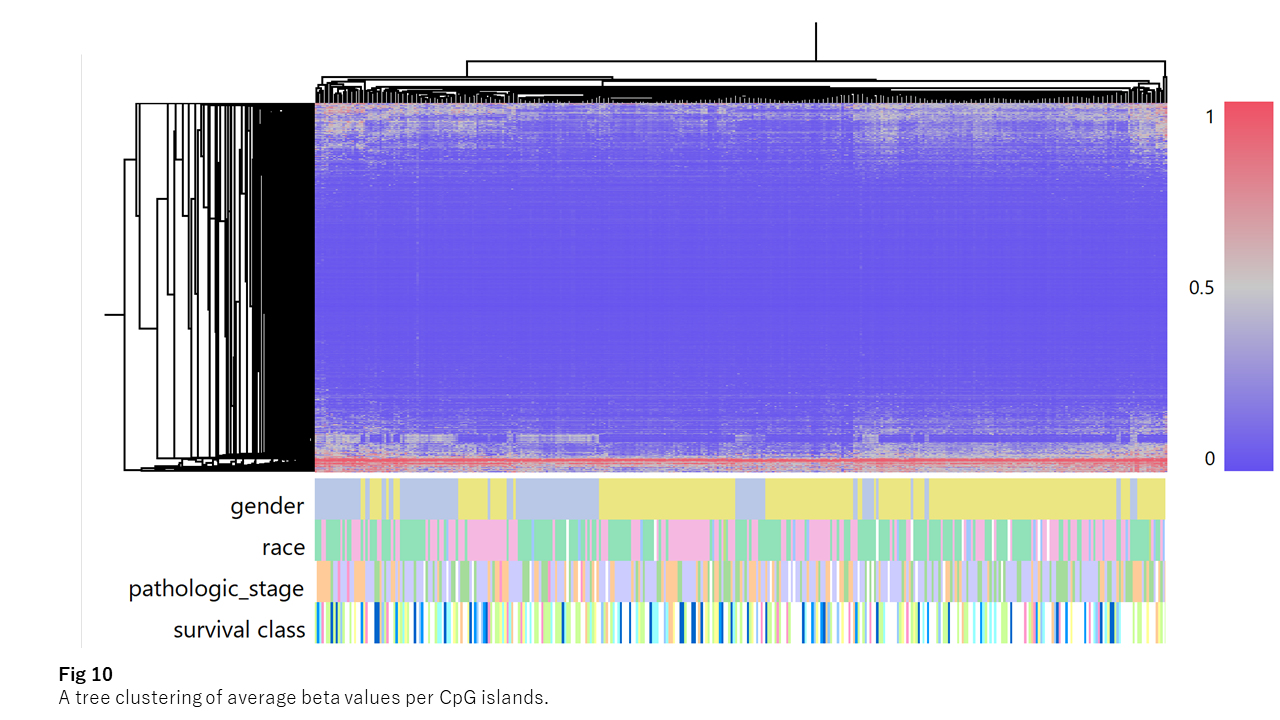
If you look at the average beta values of CpG islands over patients, most of the genes have CpG islands thoroughly maintained as unmethylated. On the other hand, a tiny number of genes have constantly highly-methylated CpG islands. The rest of about 4,000 CpG islands, whose beta values vary among the patients, possibly affect the neighboring gene’s expression level. (Fig 10)
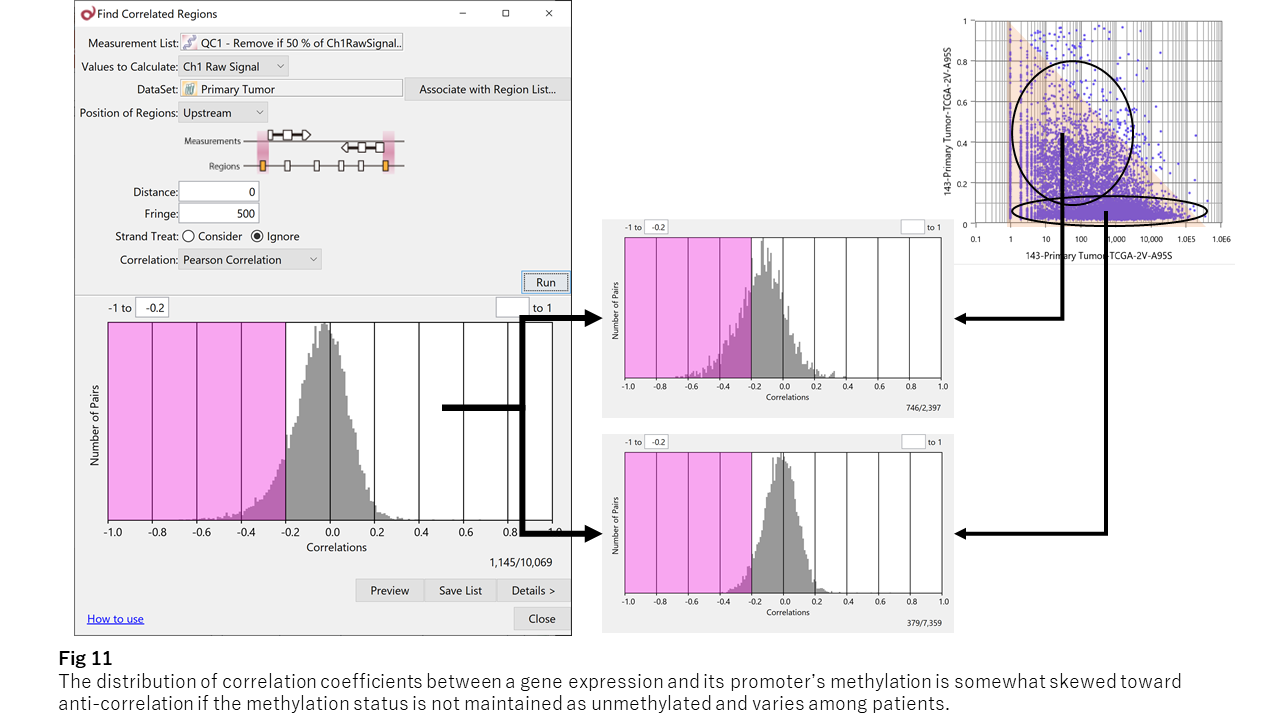
Fig 11 shows the distribution of correlation coefficients between the average beta per CpG island and the neighboring gene’s expression level. The peak lies at 0, reflecting not-correlated. And the distribution skewed toward anti-correlation, meaning the methylation of the promoter region negatively regulates neighboring gene’s expression.
Fig 10 indicates most genes have thoroughly unmethylated promoters. As expected, such genes show no correlation between the gene expression and their promoter’s methylation. Contrarily, the methylation-varying genes show "weak" negative correlations between the patterns of promoter methylation and expression.
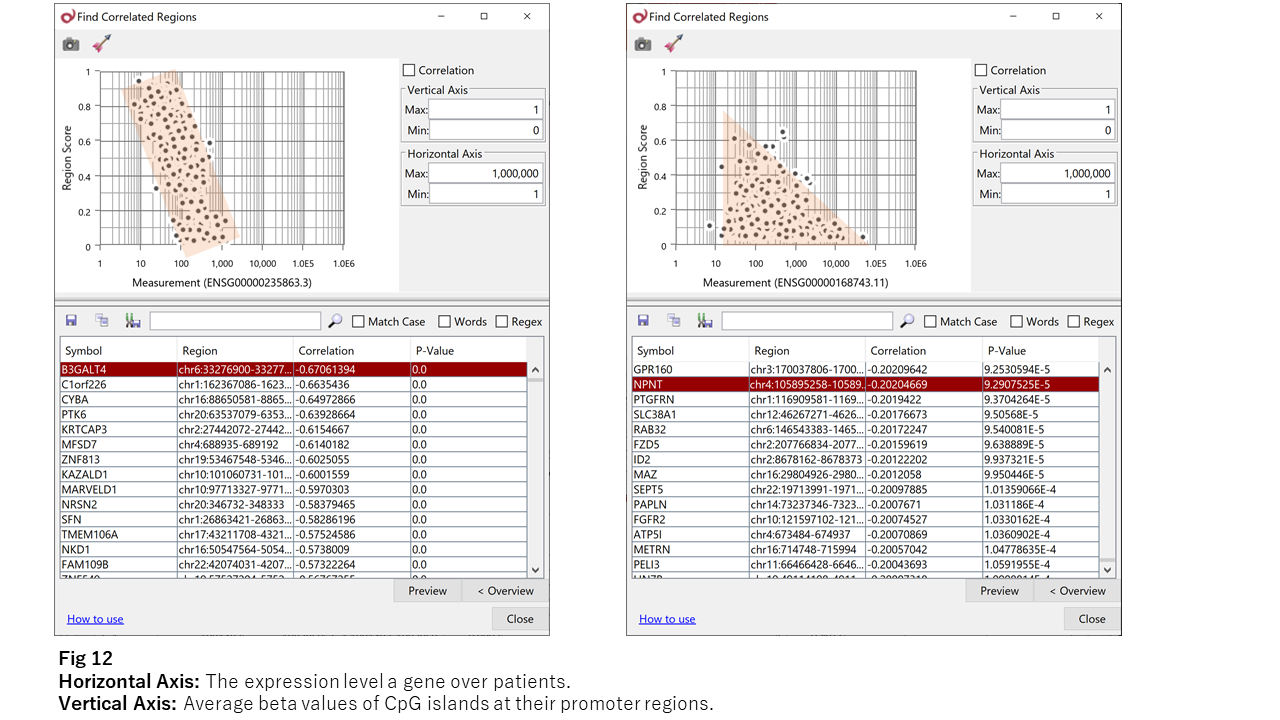
As Fig 11 indicates, only a few genes show strong anti-correlation like B3GALT4 (the left chart of Fig 12.) However, most genes show the triangle shape like NPNT (the right.) Thus, it seems that hyper-methylation can suppress the neighboring gene’s expression level, but hypo-methylation doesn’t necessarily activate. I think this is the reason for the weakness of the observed anti-correlation.
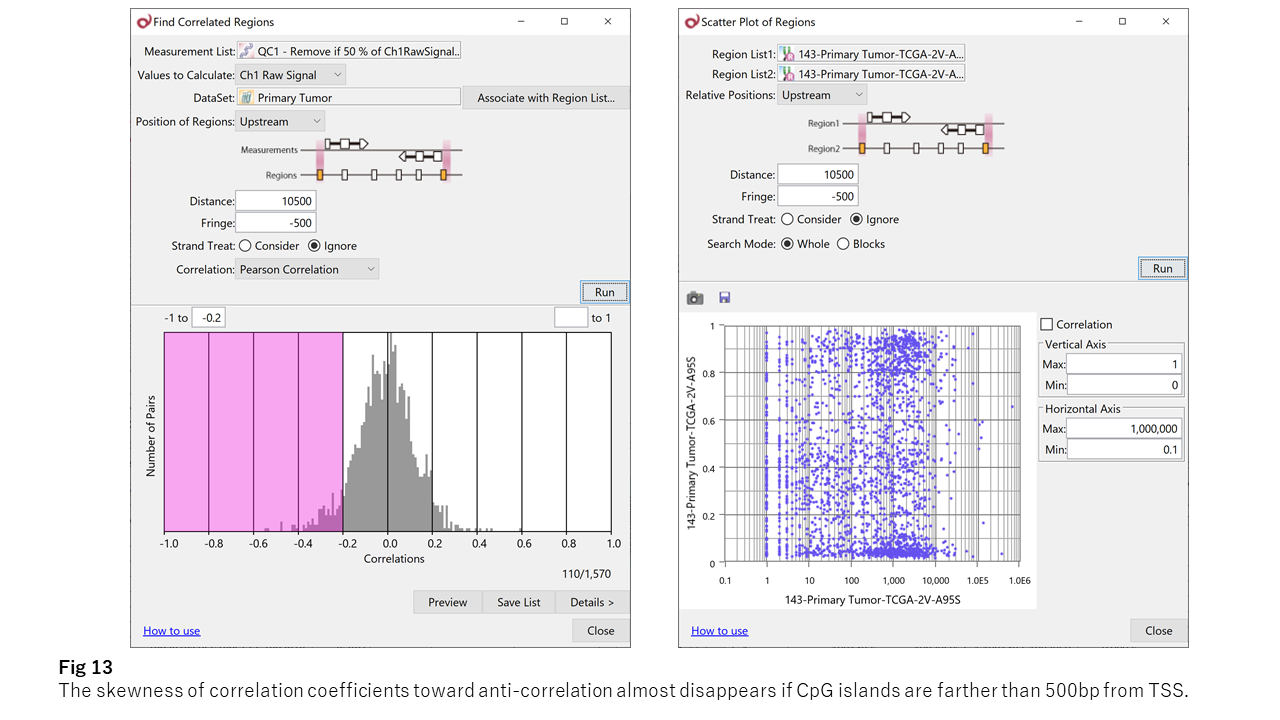
By the way, the anti-correlation is unique to the promoter (within 500bp from TSS). For example, fig 13 shows the anti-correlation between the methylation and gene expression significantly fades if the CpG island locates farther than 500bp.