RNA-Seq data are commonly transformed into metrics such as TPM or FPKM for downstream analysis.
However, are you assuming that these transformations have fully “cleaned” your data and proceeding without further validation?
The key issue is that these processing steps do not eliminate all sources of bias.
In practice, nonlinear biases often remain even after standard normalization, and these can lead to misleading conclusions in omics data analysis.
In this article, we use a real dataset (GSE159751) to demonstrate the importance of visually examining data distributions.
While there is ongoing discussion about whether TPM or FPKM should be used, the differences between them are typically minimal, and in practice, either can be used without substantially affecting the overall interpretation.
Why "Distribution Shifts" Persist After Normalization
Even after "per million" normalization, TPM/FPKM distributions often show significant shifts between samples. Furthermore, this specific dataset exhibits non-linear biases in FPKM distribution shapes (unimodal vs. bimodal). A shift to a unimodal distribution can often be a red flag for RNA degradation. If you are analyzing with Subio Platform, identifying these suspicious samples is straightforward. The key is that the analyst must understand these anomalies, decide how to handle them, and interpret the results based on that judgment. This is a crucial step often overlooked by those who neglect visualization.
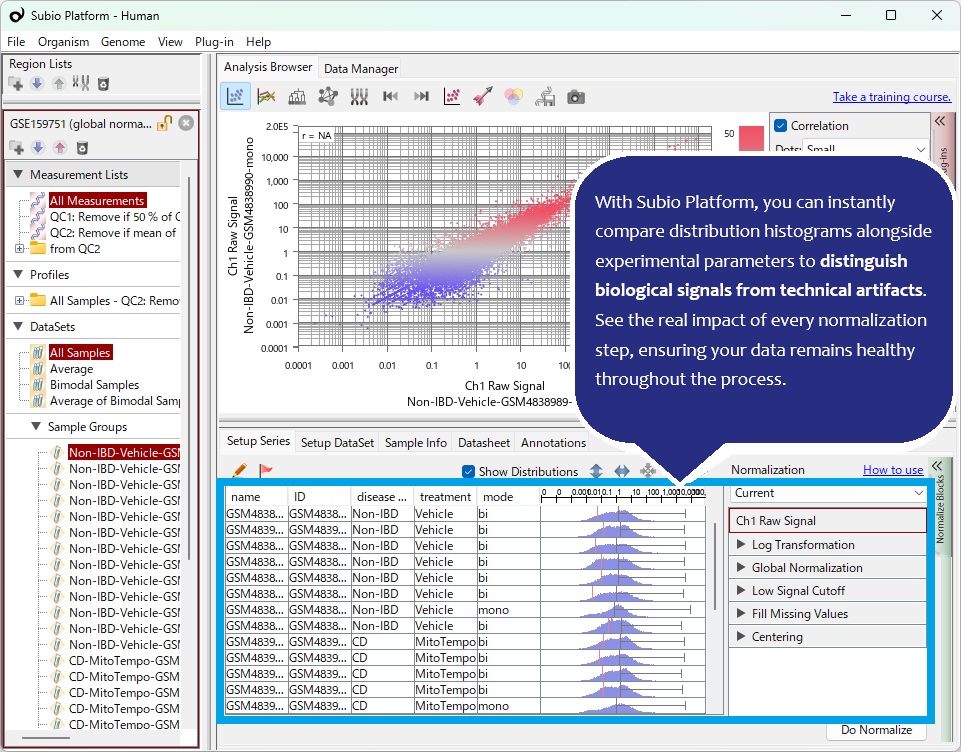
Quantile Normalization is Not a Panacea
Another vital point is evaluating the ability of "advanced" algorithms to remove non-linear bias. Here, we applied Quantile Normalization to forcibly unify the FPKM distributions. While this made the distribution shapes appear similar, it contributed nothing to the actual removal of non-linear systematic errors.
What Experimental Biologists Should Do Before Relying on Algorithms
Thanks to bioinformaticians, many algorithms are available to tackle highly complex challenges. However, wise experimental biologists must remember that they are responsible for verifying whether those tools actually work for their specific data. At this stage, generating high-quality raw data through superior experimental design and monitoring the process with accurate tools is far more effective than blindly relying on "advanced" algorithms.
When strong systematic errors enter your data, removing their influence becomes difficult or even impossible. To avoid such pitfalls, Subio believes in the importance of pre-assessment of the chosen measurement technology and experimental planning that mitigates risk.
Don't wait until the data is out to regret it. With experience handling thousands of "failed datasets," Subio can propose experimental designs that ensure success. Let us provide a professional assessment of your plan before you start your experiments. [Contact us here].
Master Analysis, Not the Tool.