“Why do samples from the same condition show larger variation than the difference between groups?”
One of the unavoidable challenges in omics data analysis is batch effect, or systematic bias. Many researchers assume that once RNA-seq data have been normalized by CPM, the samples are ready for comparison. However, depending on the state of the data, CPM normalization itself can sometimes emphasize apparent differences between samples.
In this article, we use a real dataset to show how to visually inspect what is happening in the data before relying on advanced correction algorithms.
Do Not Take Normalized Values at Face Value — Always Check the Shape of the Distribution
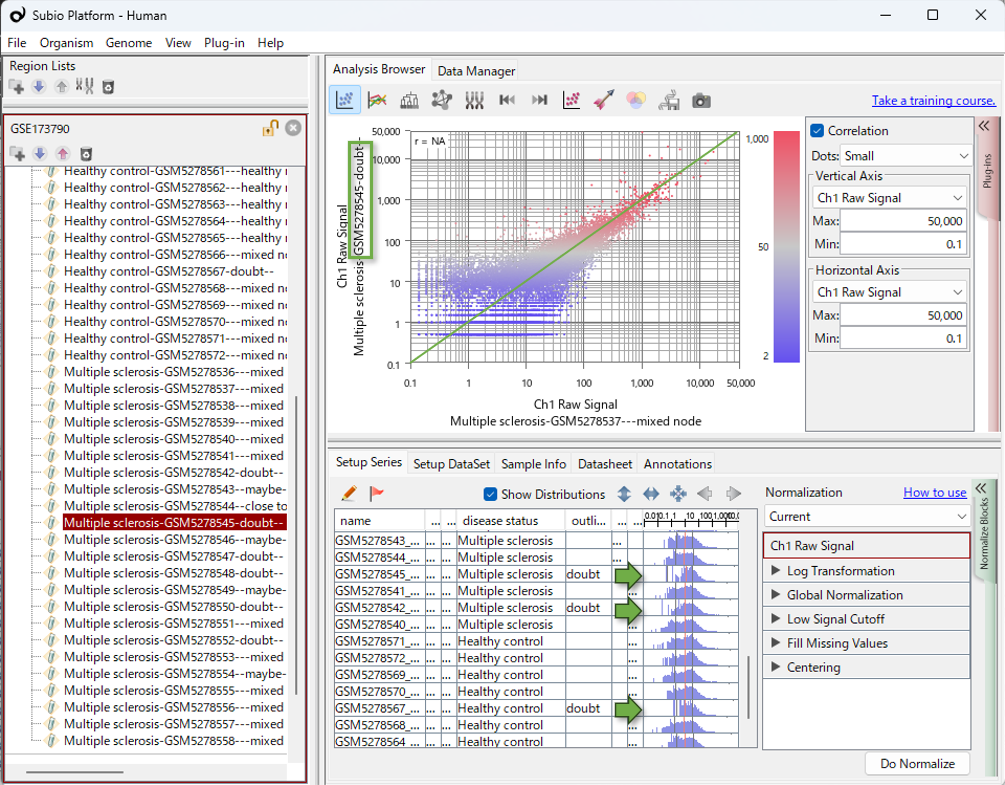
In Subio Platform, experimental parameters and histograms showing value distributions can be viewed in the same table. This makes it easier to visually check whether the data distribution appears to be related to experimental parameters or sample-level information.
In RNA-seq data, samples with extremely low read counts may show relatively elevated expression values after CPM normalization. In the histogram shown in this video, this would appear as a shift of the distribution to the right.
In this dataset, many of the samples that appear biased in PCA roughly corresponded to samples with relatively smaller read counts or FASTQ file sizes in SRA (Sequence Read Archive) . This tendency was especially clear for MS-13, MS-16 to MS-19, and HC-19.
However, a two-fold difference in FASTQ file size or read count is not unusual in RNA-seq data. Therefore, samples with smaller file sizes should not be immediately regarded as outliers. What matters is whether differences in read counts or file sizes correspond to the bias observed in PCA or to changes in the histogram distributions.
A smaller FASTQ file size suggests that fewer total reads were obtained for that sample. CPM normalization scales each sample so that the total read count is brought to the same level. As a result, in samples with fewer total reads, the same count value is adjusted to a relatively larger value. This can make expression values appear globally elevated in low-read-count samples, and the histogram distribution may appear shifted to the right.
At this point, genes with high counts may appear to shift to the right while largely preserving the overall shape of the distribution. In contrast, low-count genes are originally compressed into the left side of the Gene Count histogram. When CPM normalization shifts these values to the right, a blank region appears on the left side of the histogram. This is one possible reason why the distribution in the histogram below appears distorted.
Importantly, this does not mean that CPM normalization itself is always problematic. CPM is a simple and intuitive method, and it can be useful for checking the state of the data. Problems arise when the assumptions behind normalization do not match the state of the data. For example, differences in read depth, composition bias where reads are concentrated in specific gene groups, or data with many low-count or zero-count genes may require careful interpretation of the normalized distributions and sample-level patterns.
This issue is not limited to CPM normalization. Even normalized values such as TPM or FPKM can show apparent differences depending on sample-level data size, expression distributions, or composition bias. In RNA-seq analysis, various normalization methods are used, including TMM normalization and the size factor approach used in DESeq2, but simply changing the normalization method does not guarantee that the results are safe to interpret.
What matters is not only which normalization method was used, but whether the distributions before and after normalization, PCA, and clustering patterns have been checked, and whether the observed differences are understood in relation to the underlying sample structure.
In this dataset, based on the sample information in GEO, edgeR appears to have been used to convert Gene Count values into CPM values. edgeR is a widely used and reliable tool for RNA-seq analysis. However, performing the analysis with a reliable tool and correctly interpreting the results are two different things.
What matters is to examine whether the data entered into the tool sufficiently meet the assumptions of the analysis, and to understand what the resulting differences actually reflect. No matter how reliable the tool is, if we do not check what kind of sample structure the extracted expression differences are derived from, we may misinterpret the meaning of the results.
In this way, it is important to carefully examine whether the extracted differentially expressed genes truly reflect biological differences, or whether they include apparent differences caused by data size, normalization, or bias within the sample groups. If data analysis proceeds without visual inspection, there is a risk of misinterpreting the resulting DEG list.
Transcriptome analysis is based on the assumption that, among all genes, most expression levels remain unchanged and only a subset of genes show differential expression. This assumption is what makes normalization possible.
If the shapes of the histograms differ greatly between samples, this assumption may not hold sufficiently. Therefore, rather than proceeding with analysis based only on normalized values, it is important to examine the distribution shapes and sample-level data sizes while interpreting the results.
Data Biases Can Arise at Multiple Steps
Even after identifying samples that clearly require caution, such as those with extremely low read counts, the analytical process does not necessarily end there. Even if samples that may strongly violate the assumptions are removed and the data are visualized again, another sample structure may still appear.
In this dataset, the Control samples appear to separate into two clusters. One Control cluster shows a profile that is distant from the disease group, whereas the other Control cluster shows a profile closer to the disease group.
In such a situation, at least two possibilities need to be considered.
Possibility 1: The Pattern Reflects Biological Heterogeneity
The Control group may include samples with different biological backgrounds, and these differences may be reflected in their expression profiles. For example, age, sex, inflammatory status, medical history, or differences in cell subset composition could potentially contribute to the observed pattern.
In this case, the separation of the Control group is not simply noise. It may contain important information for interpretation. Therefore, such samples should not be removed without sufficient justification.
Possibility 2: The Pattern Reflects Experimental or Technical Bias
On the other hand, the separation of the Control group may have been caused by technical factors, such as library preparation, sequencing run, sample processing date, storage conditions, RNA quality, or other experimental factors.
In this case, the observed difference may not reflect a biological disease effect, but rather a bias introduced during data generation or processing. Whether such samples should be included in the main analysis needs to be considered carefully.
Some Questions Cannot Be Answered by Algorithms Alone
In this type of situation, simply asking which statistical method should be used is not enough. Changing the algorithm does not eliminate the underlying data structure: the Control samples still appear to separate into two groups.
What is needed here is information outside the computational results. For example, clinical background of the Control samples, sample collection conditions, library preparation date, sequencing run, RNA quality, read counts, and laboratory records may help determine whether the observed structure reflects biological differences or technical bias.
Even so, it is not always possible to obtain a clear answer to every question. In such cases, the analyst needs to clarify the assumptions being made based on the available information, and choose an analysis strategy according to those assumptions.
The important point is not to avoid making a judgment, but to make the assumptions behind the analysis explicit. Whichever interpretation is adopted, that assumption will strongly influence how the analysis results should be understood.
In omics data analysis, it is not always possible to reach conclusions through a fully mechanical workflow. It is important to visualize the state of the data, examine relevant background information when necessary, and make conscious analytical decisions.
What Small Changes May Suggest: Complementary Use of Bulk and Single-Cell Analysis
Suppose we decide that some of the Control samples that appear far from the disease group are more likely to reflect technical bias than biological differences. In that case, the expression differences observed between the remaining Control samples and the disease group may become very small.
In bulk RNA-seq, many different cells are measured together. If important changes occur only in a small subset of cells, the overall expression difference may appear small. Therefore, it is natural to consider whether the data should be examined by single-cell RNA-seq, or whether the target cells should be further enriched by cell sorting and the experiment repeated.
At the same time, there is another important point to keep in mind. Single-cell analysis is extremely useful for examining differences among cell populations, but its data characteristics differ from those of bulk RNA-seq. It can be strongly affected by dropout, measurement noise, cell number, and preprocessing conditions.
Therefore, before chasing subtle signals with another method, it may be useful to first examine whether small but consistent expression differences can be observed in bulk RNA-seq, where measurement stability is often higher.
The important question is not “bulk or single-cell?” These approaches can be used in a complementary way. Bulk RNA-seq can be used to examine the overall picture and reproducible changes, while single-cell analysis or cell fractionation experiments can be used when needed to deepen the interpretation at the cell population level.
What Is the Role of the Analyst?
The discussion above describes the process of how I interpreted this dataset while examining it. It is not intended to claim that this is the only correct answer.
What I want to emphasize is the importance of considering multiple possibilities while looking at the data and clarifying the points at which interpretation may diverge. When it is not possible to narrow the explanation down to a single possibility, the analyst needs to organize the results based on each possible scenario and present them in a way that can support discussion with team leaders, experimental researchers, clinicians, pathologists, and others who understand the background of the data.
Mastering advanced statistical methods alone is not the essence of omics data analysis. Visualizing the data, checking assumptions, and carefully explaining the characteristics of the data and the analysis results are also important parts of the analyst’s work.
What It Means to Learn Data Analysis
Learning data analysis means gaining experience by working with many different datasets, visualizing them, forming hypotheses, checking those hypotheses, and making analytical decisions. Naturally, this cannot be mastered overnight. It requires time, persistence, and repeated practice.
How Subio Supports Your Growth
Subio provides practical learning through online training, where we work together with customers using their own data.
Although the pace varies from person to person, many users can steadily build their analytical thinking and decision-making skills through approximately three to six one-on-one sessions over the course of about one year, using their own data as the learning material.
For those who need results quickly, or who first want their data to be examined from an expert perspective, we recommend our data analysis service.
Subio’s data analysis service is not simply a contract analysis service that delivers a results report. It is designed to help customers understand the characteristics of their data and the analysis steps, and to provide the information needed for customers themselves to make decisions and draw conclusions.
If you are interested, please feel free to contact us.
What you need to learn is not only commands or how to operate tools.
What you truly need to learn is data analysis itself.
Related Topics
- Do Not Trust TPM, FPKM, or RPKM Too Much: Systematic Biases in RNA-Seq Data Are Not Always Removed
- Before Relying on Plug-ins: Why the Basic Functions of Subio Platform Are Your Best Analysis Partner
- Omics Experiments Cannot Escape the Problem of Nonlinear Bias
- An Efficient Way to Learn Omics Data Analysis